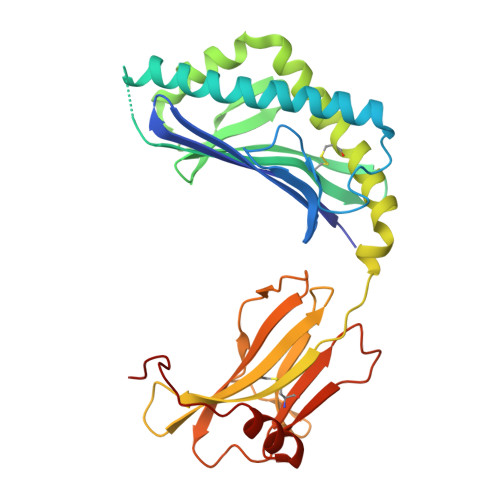
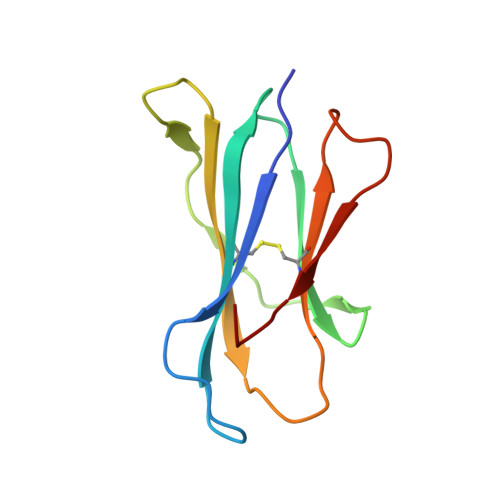
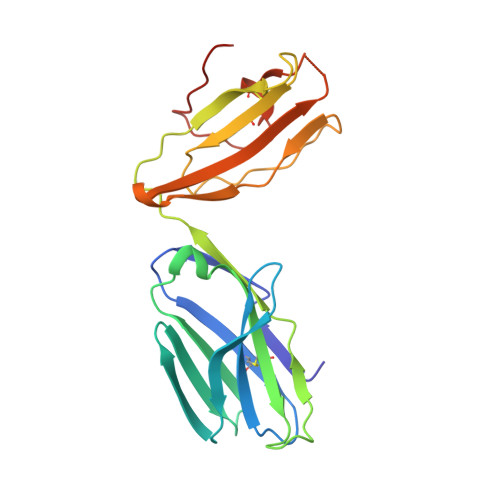
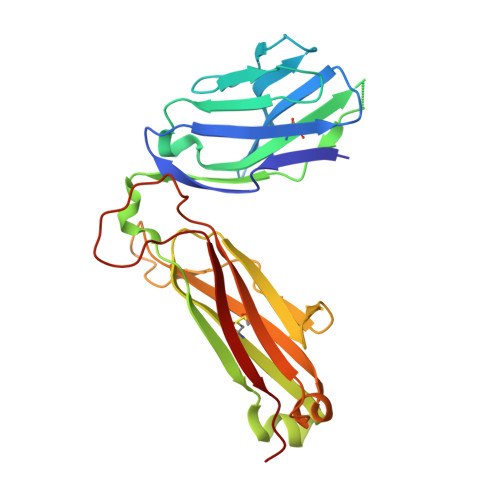
A molecular basis for the exquisite CD1d-restricted antigen specificity and functional responses of natural killer T cells
Wun, K.S., Cameron, G., Patel, O., Pang, S.S., Pellicci, D.G., Sullivan, L.C., Keshipeddy, S., Young, M.H., Uldrich, A.P., Thakur, M.S., Richardson, S.K., Howell, A.R., Illarionov, P.A., Brooks, A.G., Besra, G.S., McCluskey, J., Gapin, L., Porcelli, S.A., Godfrey, D.I., Rossjohn, J.(2011) Immunity 34: 327-339
- PubMed: 21376639 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.immuni.2011.02.001
- Primary Citation Related Structures:
3ARB, 3ARD, 3ARE, 3ARF, 3ARG - PubMed Abstract:
Natural killer T (NKT) cells respond to a variety of CD1d-restricted antigens (Ags), although the basis for Ag discrimination by the NKT cell receptor (TCR) is unclear. Here we have described NKT TCR fine specificity against several closely related Ags, termed altered glycolipid ligands (AGLs), which differentially stimulate NKT cells. The structures of five ternary complexes all revealed similar docking. Acyl chain modifications did not affect the interaction, but reduced NKT cell proliferation, indicating an affect on Ag processing or presentation. Conversely, truncation of the phytosphingosine chain caused an induced fit mode of TCR binding that affected TCR affinity. Modifications in the glycosyl head group had a direct impact on the TCR interaction and associated cellular response, with ligand potency reflecting the t(1/2) life of the interaction. Accordingly, we have provided a molecular basis for understanding how modifications in AGLs can result in striking alterations in the cellular response of NKT cells.
- The Protein Crystallography Unit, ARC Centre of Excellence in Structural and Functional Microbial Genomics, Department of Biochemistry and Molecular Biology, School of Biomedical Sciences, Monash University, Clayton, Victoria 3800, Australia.
Organizational Affiliation: