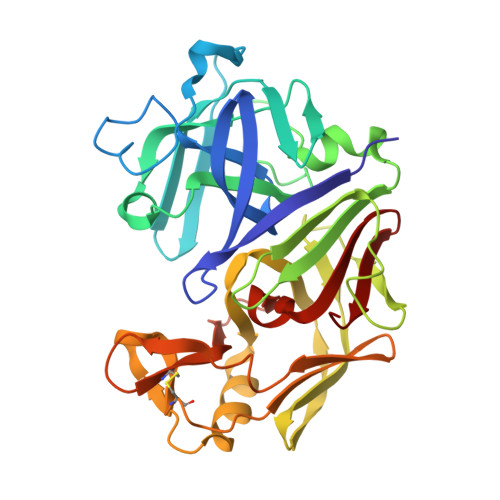
Structure and refinement of penicillopepsin at 1.8 A resolution.
James, M.N., Sielecki, A.R.(1983) J Mol Biology 163: 299-361
- PubMed: 6341600 Search on PubMed
- DOI: https://doi.org/10.1016/0022-2836(83)90008-6
- Primary Citation Related Structures:
3APP