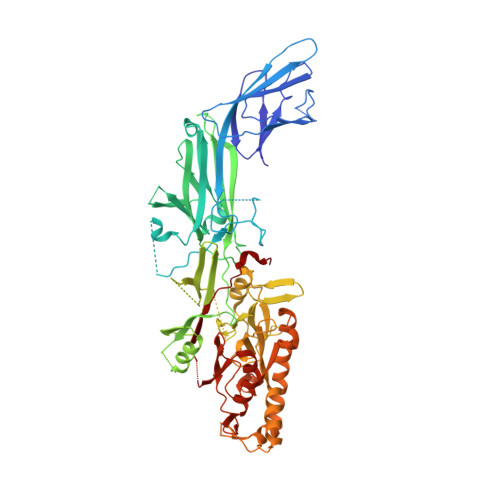
Structural and biochemical analyses of the human PAD4 variant encoded by a functional haplotype gene
Horikoshi, N., Tachiwana, H., Saito, K., Osakabe, A., Sato, M., Yamada, M., Akashi, S., Nishimura, Y., Kagawa, W., Kurumizaka, H.(2011) Acta Crystallogr D Biol Crystallogr 67: 112-118
- PubMed: 21245532 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444910051711
- Primary Citation Related Structures:
3APM, 3APN - PubMed Abstract:
PAD4 is a peptidylarginine deiminase that catalyzes citrullination, a type of post-translational modification. In this reaction, arginine residues in proteins are converted to citrulline. PAD4 promotes the deimination of arginine residues in histones and may regulate transcription in the context of the chromatin. Single-nucleotide polymorphisms (SNP) in the gene encoding PAD4 identified it as one of the genes associated with susceptibility to rheumatoid arthritis. The PAD4 SNP involve three amino-acid substitutions: Ser55 to Gly, Ala82 to Val and Ala112 to Gly. Autoantibodies for improperly citrullinated proteins have been found in rheumatoid arthritis patients, suggesting that the PAD4(SNP) mRNA is more stable than the conventional PAD4 mRNA and/or the PAD4(SNP) protein possesses a higher citrullination activity than the PAD4 protein. In order to study the effects of the three amino-acid substitutions found in PAD4(SNP), the crystal structure of PAD4(SNP) was determined and it was found that the amino-acid substitutions in PAD4(SNP) only induced conformational changes within the N-terminal domain, not in the active centre for citrullination located in the C-terminal domain. Biochemical analyses also suggested that the citrullination activity of PAD4(SNP) may not substantially differ from that of conventional PAD4. These structural and biochemical findings suggested that the improper protein citrullination found in rheumatoid arthritis patients is not caused by defects in the citrullination activity of PAD4(SNP) but by other reasons such as enhanced PAD4(SNP) mRNA stability.
- Laboratory of Structural Biology, Graduate School of Advanced Science and Engineering, Waseda University, 2-2 Wakamatsu-cho, Shinjyuku-ku, Tokyo 162-8480, Japan.
Organizational Affiliation: