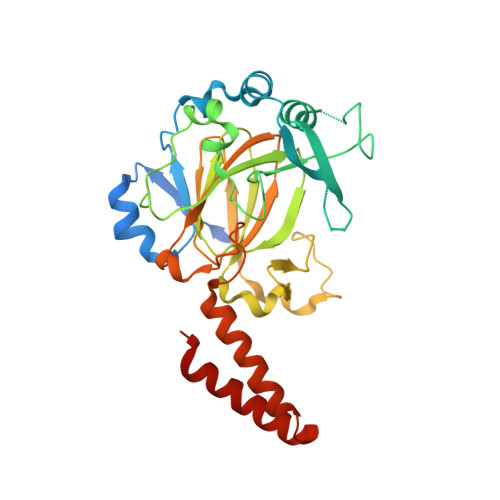
Crystal structure of a novel JmjC-domain-containing protein, TYW5, involved in tRNA modification.
Kato, M., Araiso, Y., Noma, A., Nagao, A., Suzuki, T., Ishitani, R., Nureki, O.(2011) Nucleic Acids Res 39: 1576-1585
- PubMed: 20972222 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkq919
- Primary Citation Related Structures:
3AL5, 3AL6 - PubMed Abstract:
Wybutosine (yW) is a hypermodified nucleoside found in position 37 of tRNA(Phe), and is essential for correct phenylalanine codon translation. yW derivatives widely exist in eukaryotes and archaea, and their chemical structures have many species-specific variations. Among them, its hydroxylated derivative, hydroxywybutosine (OHyW), is found in eukaryotes including human, but the modification mechanism remains unknown. Recently, we identified a novel Jumonji C (JmjC)-domain-containing protein, TYW5 (tRNA yW-synthesizing enzyme 5), which forms the OHyW nucleoside by carbon hydroxylation, using Fe(II) ion and 2-oxoglutarate (2-OG) as cofactors. In this work, we present the crystal structures of human TYW5 (hTYW5) in the free and complex forms with 2-OG and Ni(II) ion at 2.5 and 2.8 Å resolutions, respectively. The structure revealed that the catalytic domain consists of a β-jellyroll fold, a hallmark of the JmjC domains and other Fe(II)/2-OG oxygenases. hTYW5 forms a homodimer through C-terminal helix bundle formation, thereby presenting a large, positively charged patch involved in tRNA binding. A comparison with the structures of other JmjC-domain-containing proteins suggested a mechanism for substrate nucleotide recognition. Functional analyses of structure-based mutants revealed the essential Arg residues participating in tRNA recognition by TYW5. These findings extend the repertoire of the tRNA modification enzyme into the Fe(II)/2-OG oxygenase superfamily.
- Department of Biophysics and Biochemistry, Graduate School of Science, The University of Tokyo, 2-11-16 Yayoi, Bunkyo-ku, Tokyo 113-0032, Japan.
Organizational Affiliation: