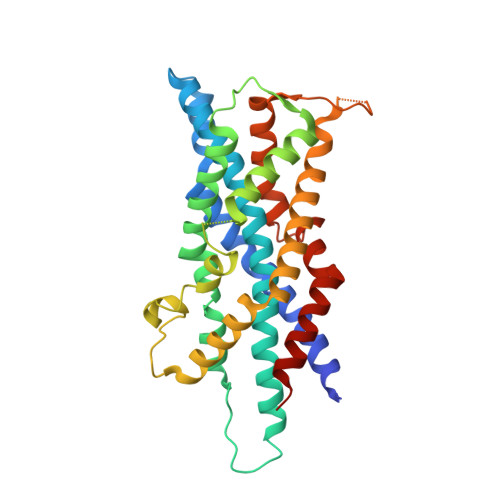
Structural basis for docking of peroxisomal membrane protein carrier Pex19p onto its receptor Pex3p
Sato, Y., Shibata, H., Nakatsu, T., Nakano, H., Kashiwayama, Y., Imanaka, T., Kato, H.(2010) EMBO J 29: 4083-4093
- PubMed: 21102411 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/emboj.2010.293
- Primary Citation Related Structures:
3AJB - PubMed Abstract:
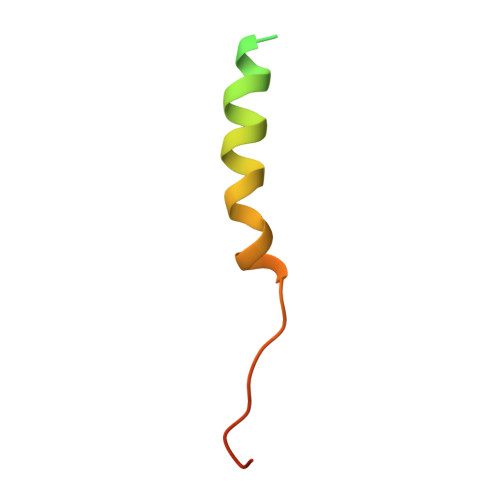
Peroxisomes require peroxin (Pex) proteins for their biogenesis. The interaction between Pex3p, which resides on the peroxisomal membrane, and Pex19p, which resides in the cytosol, is crucial for peroxisome formation and the post-translational targeting of peroxisomal membrane proteins (PMPs). It is not known how Pex3p promotes the specific interaction with Pex19p for the purpose of PMP translocation. Here, we present the three-dimensional structure of the complex between a cytosolic domain of Pex3p and the binding-region peptide of Pex19p. The overall shape of Pex3p is a prolate spheroid with a novel fold, the 'twisted six-helix bundle.' The Pex19p-binding site is at an apex of the Pex3p spheroid. A 16-residue region of the Pex19p peptide forms an α-helix and makes a contact with Pex3p; this helix is disordered in the unbound state. The Pex19p peptide contains a characteristic motif, consisting of the leucine triad (Leu18, Leu21, Leu22), and Phe29, which are critical for the Pex3p binding and peroxisome biogenesis.
- Department of Structural Biology, Graduate School of Pharmaceutical Sciences, Kyoto University, Kyoto, Japan.
Organizational Affiliation: