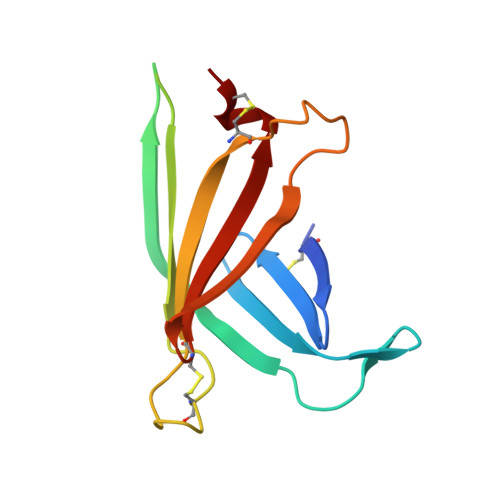
Structural Basis for Oligosaccharide Recognition of Misfolded Glycoproteins by OS-9 in ER-Associated Degradation
Satoh, T., Chen, Y., Hu, D., Hanashima, S., Yamamoto, K., Yamaguchi, Y.(2010) Mol Cell 40: 905-916
- PubMed: 21172656 Search on PubMed
- DOI: https://doi.org/10.1016/j.molcel.2010.11.017
- Primary Citation Related Structures:
3AIH - PubMed Abstract:
Misfolded glycoproteins are translocated from endoplasmic reticulum (ER) into the cytosol for proteasome-mediated degradation. A mannose-6-phosphate receptor homology (MRH) domain is commonly identified in a variety of proteins and, in the case of OS-9 and XTP3-B, is involved in glycoprotein ER-associated degradation (ERAD). Trimming of outermost α1,2-linked mannose on C-arm of high-mannose-type glycan and binding of processed α1,6-linked mannosyl residues by the MRH domain are critical steps in guiding misfolded glycoproteins to enter ERAD. Here we report the crystal structure of a human OS-9 MRH domain (OS-9(MRH)) complexed with α3,α6-mannopentaose. The OS-9(MRH) has a flattened β-barrel structure with a characteristic P-type lectin fold and possesses distinctive double tryptophan residues in the oligosaccharide-binding site. Our crystallographic result in conjunction with nuclear magnetic resonance (NMR) spectroscopic and biochemical results provides structural insights into the mechanism whereby OS-9 specifically recognizes Manα1,6Manα1,6Man residues on the processed C-arm through the continuous double tryptophan (WW) motif.
- Structural Glycobiology Team, RIKEN Advanced Science Institute, Saitama 351-0198, Japan.
Organizational Affiliation: