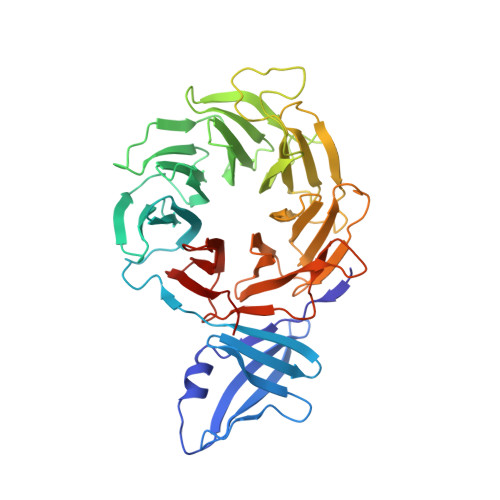
Crystal structure of yeast Rpn14, a chaperone of the 19S regulatory particle of the proteasome
Kim, S., Saeki, Y., Fukunaga, K., Suzuki, A., Takagi, K., Yamane, T., Tanaka, K., Mizushima, T., Kato, K.(2010) J Biological Chem 285: 15159-15166
- PubMed: 20236927 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M110.104042
- Primary Citation Related Structures:
3ACP - PubMed Abstract:
The ubiquitin-proteasome pathway is a major proteolytic system in eukaryotic cells and regulates various cellular processes. The 26 S proteasome, the central enzyme of this pathway, consists of a proteolytic core particle and two 19 S regulatory particles (RPs) composed of ATPase (Rpt) and non-ATPase (Rpn) subunits. Growing evidence indicates that proteasome assembly is assisted by a variety of chaperones. In particular, it has been reported recently that Nas2, Nas6, Rpn14, and Hsm3 bind specific Rpt subunits, thereby contributing to the formation of 19 S RP. Rpn14 transiently binds to the C-terminal domain of the Rpt6 subunit (Rpt6-C) during maturation of the ATPase ring of 19 S RP. In this study, we determined the crystal structure of yeast Rpn14 at 2.0 A resolution, which revealed that this chaperone consists of a unique N-terminal domain with unknown function and a C-terminal domain assuming a canonical seven-bladed beta-propeller fold. The Rpt6-binding site on Rpn14 was predicted based on structural comparison with the complex formed between Nas6 and Rpt3-C. The top face of Rpn14 exhibits a highly acidic surface area, whereas the putative interacting surface of Rpt6-C is basic. By inspection of structural data along with genetic and biochemical data, we determined the specific residues of Rpn14 and Rpt6 for complementary charge interactions that are required for 19 S RP assembly.
- Graduate School of Pharmaceutical Sciences, Nagoya City University, 3-1 Tanabe-dori, Mizuho-ku, Nagoya 467-8603.
Organizational Affiliation: