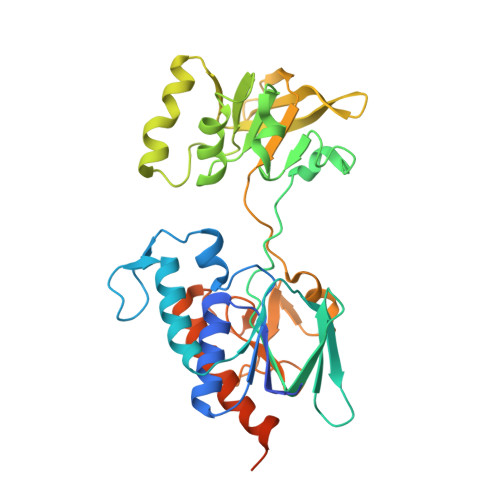
Asymmetric dimeric structure of ferredoxin-NAD(P)+ oxidoreductase from the green sulfur bacterium Chlorobaculum tepidum: implications for binding ferredoxin and NADP+
Muraki, N., Seo, D., Shiba, T., Sakurai, T., Kurisu, G.(2010) J Mol Biology 401: 403-414
- PubMed: 20600130 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2010.06.024
- Primary Citation Related Structures:
3AB1 - PubMed Abstract:
Ferredoxin-NAD(P)(+) oxidoreductase (FNR) catalyzes the reduction of NAD(P)(+) to NAD(P)H with the reduced ferredoxin (Fd) during the final step of the photosynthetic electron transport chain. FNR from the green sulfur bacterium Chlorobaculum tepidum is functionally analogous to plant-type FNR but shares a structural homology to NADPH-dependent thioredoxin reductase (TrxR). Here, we report the crystal structure of C. tepidum FNR to 2.4 A resolution, which reveals a unique structure-function relationship. C. tepidum FNR consists of two functional domains for binding FAD and NAD(P)H that form a homodimer in which the domains are arranged asymmetrically. One NAD(P)H domain is present as the open form, the other with the equivalent NAD(P)H domain as the relatively closed form. We used site-directed mutagenesis on the hinge region connecting the two domains in order to investigate the importance of the flexible hinge. The asymmetry of the NAD(P)H domain and the comparison with TrxR suggested that the hinge motion might be involved in pyridine nucleotide binding and binding of Fd. Surprisingly, the crystal structure revealed an additional C-terminal sub-domain that tethers one protomer and interacts with the other protomer by pi-pi stacking of Phe337 and the isoalloxazine ring of FAD. The position of this stacking Phe337 is almost identical with both of the conserved C-terminal Tyr residues of plant-type FNR and the active site dithiol of TrxR, implying a unique structural basis for enzymatic reaction of C. tepidum FNR.
- Department of Life Sciences, University of Tokyo, Komaba, Meguro-ku, Tokyo 153-8902, Japan.
Organizational Affiliation: