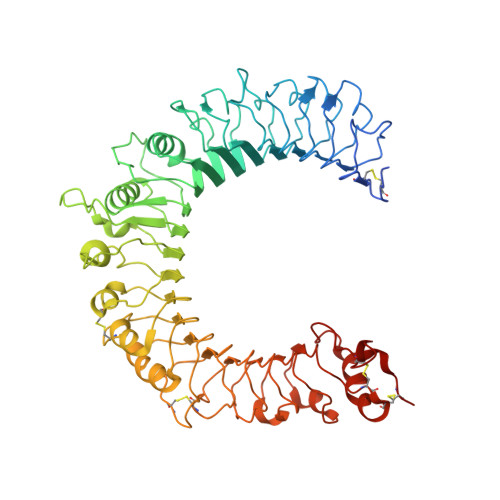
Recognition of lipopeptide patterns by Toll-like receptor 2-Toll-like receptor 6 heterodimer
Kang, J.Y., Nan, X., Jin, M.S., Youn, S.-J., Ryu, Y.H., Mah, S., Han, S.H., Lee, H., Paik, S.-G., Lee, J.-O.(2009) Immunity 31: 873-884
- PubMed: 19931471 Search on PubMed
- DOI: https://doi.org/10.1016/j.immuni.2009.09.018
- Primary Citation Related Structures:
3A79, 3A7B, 3A7C - PubMed Abstract:
Toll-like receptor 2 (TLR2) initiates potent immune responses by recognizing diacylated and triacylated lipopeptides. Its ligand specificity is controlled by whether it heterodimerizes with TLR1 or TLR6. We have determined the crystal structures of TLR2-TLR6-diacylated lipopeptide, TLR2-lipoteichoic acid, and TLR2-PE-DTPA complexes. PE-DTPA, 1,2-dimyristoyl-sn-glycero-3-phosphoethanolamine-N-diethylenetriaminepentaacetic acid, is a synthetic phospholipid derivative. Two major factors contribute to the ligand specificity of TLR2-TLR1 or TLR2-TLR6 heterodimers. First, the lipid channel of TLR6 is blocked by two phenylalanines. Simultaneous mutation of these phenylalanines made TLR2-TLR6 fully responsive not only to diacylated but also to triacylated lipopeptides. Second, the hydrophobic dimerization interface of TLR2-TLR6 is increased by 80%, which compensates for the lack of amide lipid interaction between the lipopeptide and TLR2-TLR6. The structures of the TLR2-lipoteichoic acid and the TLR2-PE-DTPA complexes demonstrate that a precise interaction pattern of the head group is essential for a robust immune response by TLR2 heterodimers.
- Department of Chemistry, KAIST, Daejon, 305-701, Korea.
Organizational Affiliation: