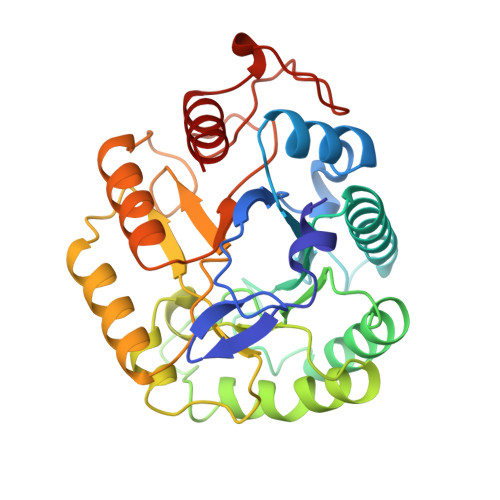
Snapshots along an enzymatic reaction coordinate: analysis of a retaining beta-glycoside hydrolase.
Davies, G.J., Mackenzie, L., Varrot, A., Dauter, M., Brzozowski, A.M., Schulein, M., Withers, S.G.(1998) Biochemistry 37: 11707-11713
- PubMed: 9718293 Search on PubMed
- DOI: https://doi.org/10.1021/bi981315i
- Primary Citation Related Structures:
3A3H, 4A3H, 5A3H, 6A3H, 7A3H - PubMed Abstract:
The enzymatic hydrolysis of O-glycosidic linkages is one of the most diverse and widespread reactions in nature and involves a classic "textbook" enzyme mechanism. A multidisciplinary analysis of a beta-glycoside hydrolase, the Cel5A from Bacillus agaradhaerens, is presented in which the structures of each of the native, substrate, covalent-intermediate, and product complexes have been determined and their interconversions analyzed kinetically, providing unprecedented insights into the mechanism of this enzyme class. Substrate is bound in a distorted 1S3 skew-boat conformation, thereby presenting the anomeric carbon appropriately for nucleophilic attack as well as satisfying the stereoelectronic requirements for an incipient oxocarbenium ion. Leaving group departure results in the trapping of a covalent alpha-glycosyl-enzyme intermediate in which the sugar adopts an undistorted 4C1 conformation. Finally, hydrolysis of this intermediate yields a product complex in which the sugar is bound in a partially disordered mode, consistent with unfavorable interactions and low product affinity.
- Department of Chemistry, University of York, Heslington, U.K. davies@yorvic.york.ac.uk
Organizational Affiliation: