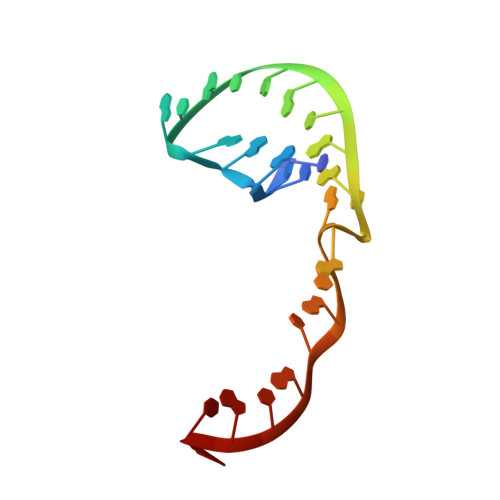
The Structure of an RNA Pseudoknot Shows 3D Domain Swapping
Lietzke, S.E., Barnes, C.L., Malone, V.F., Jones, J.T., Kundrot, C.E.(1998) Structure, Motion, Interaction And Expression Of Biological Macromolecules, The Proceedings Of The Tenth Conversation Held At The University-suny, Albany Ny, June 17-21, 1997 10: 91-101