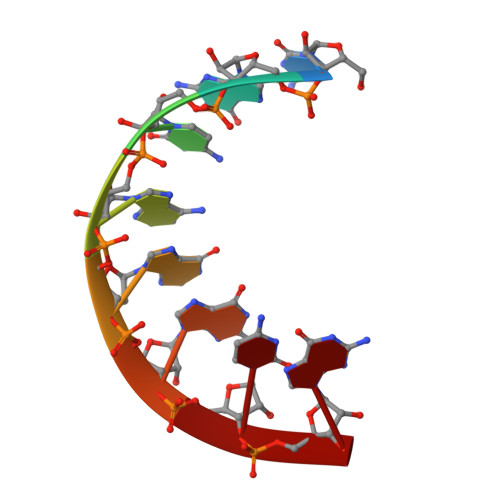
The crystal structure of an RNA oligomer incorporating tandem adenosine-inosine mismatches.
Carter, R.J., Baeyens, K.J., SantaLucia, J., Turner, D.H., Holbrook, S.R.(1997) Nucleic Acids Res 25: 4117-4122
- PubMed: 9321667 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/25.20.4117
- Primary Citation Related Structures:
333D - PubMed Abstract:
The X-ray crystallographic structure of the RNA duplex [r(CGCAIGCG)]2 has been refined to 2.5 A. It shows a symmetric internal loop of two non-Watson-Crick base pairs which form in the middle of the duplex. The tandem A-I/I-A pairs are related by a crystallographic two-fold axis. Both A(anti)-I(anti) mismatches are in a head-to-head conformation forming hydrogen bonds using the Watson-Crick positions. The octamer duplexes stack above one another in the cell forming a pseudo-infinite helix throughout the crystal. A hydrated calcium ion bridges between the 3'-terminal of one molecule and the backbone of another. The tandem A-I mismatches are incorporated with only minor distortion to the backbone. This is in contrast to the large helical perturbations often produced by sheared G-A pairs in RNA oligonucleotides.
- Structural Biology Division, Lawrence Berkeley Laboratory, Berkeley, CA 94720, USA.
Organizational Affiliation: