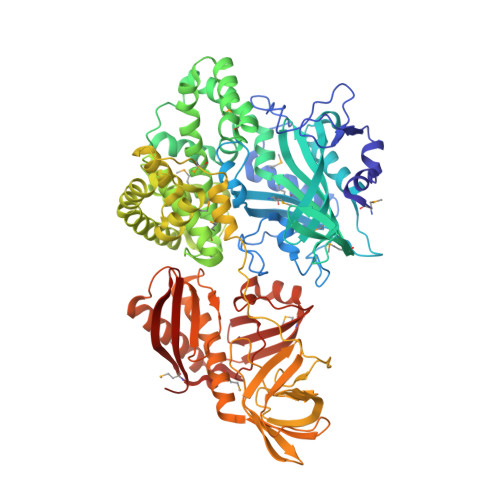
Unique protein architecture of alanyl-tRNA synthetase for aminoacylation, editing, and dimerization.
Naganuma, M., Sekine, S., Fukunaga, R., Yokoyama, S.(2009) Proc Natl Acad Sci U S A 106: 8489-8494
- PubMed: 19423669 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0901572106
- Primary Citation Related Structures:
2ZTG, 2ZVF - PubMed Abstract:
Alanyl-tRNA synthetase (AlaRS) specifically recognizes the major identity determinant, the G3:U70 base pair, in the acceptor stem of tRNA(Ala) by both the tRNA-recognition and editing domains. In this study, we solved the crystal structures of 2 halves of Archaeoglobus fulgidus AlaRS: AlaRS-DeltaC, comprising the aminoacylation, tRNA-recognition, and editing domains, and AlaRS-C, comprising the dimerization domain. The aminoacylation/tRNA-recognition domains contain an insertion incompatible with the class-specific tRNA-binding mode. The editing domain is fixed tightly via hydrophobic interactions to the aminoacylation/tRNA-recognition domains, on the side opposite from that in threonyl-tRNA synthetase. A groove formed between the aminoacylation/tRNA-recognition domains and the editing domain appears to be an alternative tRNA-binding site, which might be used for the aminoacylation and/or editing reactions. Actually, the amino acid residues required for the G3:U70 recognition are mapped in this groove. The dimerization domain consists of helical and globular subdomains. The helical subdomain mediates dimerization by forming a helix-loop-helix zipper. The globular subdomain, which is important for the aminoacylation and editing activities, has a positively-charged face suitable for tRNA binding.
- Department of Biophysics and Biochemistry, Graduate School of Science, University of Tokyo, 7-3-1 Hongo, Bunkyo-ku, Tokyo 113-0033, Japan.
Organizational Affiliation: