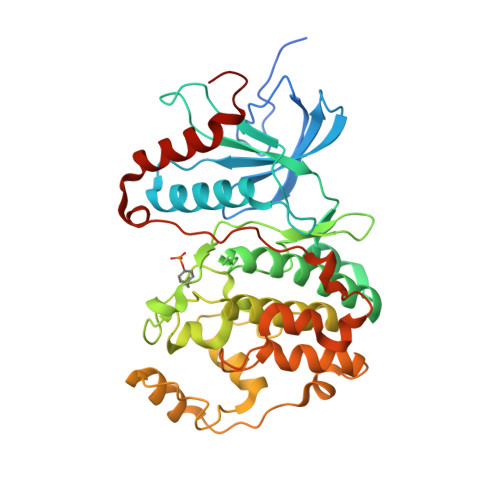
Crystal structure of human mono-phosphorylated ERK1 at Tyr204
Kinoshita, T., Yoshida, I., Nakae, S., Okita, K., Gouda, M., Matsubara, M., Yokota, K., Ishiguro, H., Tada, T.(2008) Biochem Biophys Res Commun 377: 1123-1127
- PubMed: 18983981 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2008.10.127
- Primary Citation Related Structures:
2ZOQ - PubMed Abstract:
Extracellular signal-regulated kinase (ERK) is a member of the MAP kinase family, and can regulate several cellular responses. The isoforms ERK1 and ERK2 have markedly similar amino acid sequences, but exhibit distinctive physiological functions. As well as ERK2, ERK1 was auto- and mono-phosphorylated at Tyr204 in the activation loop during Escherichia coli production, resulting in basal level activity, approximately 500-fold less compared with fully-active ERK1 dual-phosphorylated at Thr202 and Tyr204. Crystal structure demonstrated that the mono-phosphorylated ERK1 kinase possessed a novel conformation distinguishable from the un-phosphorylated (inactive) and the dual-phosphorylated (full-active) forms. The characteristic structural features in both the C-helix and the activation loop likely contribute to the basal activity of the mono-phosphorylated ERK1. The structural dissection of ERK1 compared to ERK2 suggests that the structural differences in the D-motif binding site and in the backside binding site are putative targets for development of selective ERK1/ERK2 inhibitors.
- Graduate School of Science, Osaka Prefecture University, 1-1 Gakuen-cho, Naka-ku, Sakai, Osaka 599-8531, Japan. kinotk@b.s.osakafu-u.ac.jp
Organizational Affiliation: