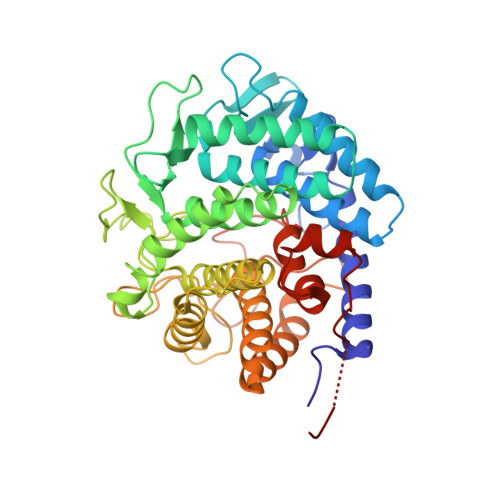
Crystal structure of YihS in complex with D-mannose: structural annotation of Escherichia coli and Salmonella enterica yihS-encoded proteins to an aldose-ketose isomerase
Itoh, T., Mikami, B., Hashimoto, W., Murata, K.(2008) J Mol Biology 377: 1443-1459
- PubMed: 18328504 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2008.01.090
- Primary Citation Related Structures:
2RGK, 2ZBL - PubMed Abstract:
The three-dimensional structure of a Salmonella enterica hypothetical protein YihS is significantly similar to that of N-acyl-D-glucosamine 2-epimerase (AGE) with respect to a common scaffold, an alpha6/alpha6-barrel, although the function of YihS remains to be clarified. To identify the function of YihS, Escherichia coli and S. enterica YihS proteins were overexpressed in E. coli, purified, and characterized. Both proteins were found to show no AGE activity but showed cofactor-independent aldose-ketose isomerase activity involved in the interconversion of monosaccharides, mannose, fructose, and glucose, or lyxose and xylulose. In order to clarify the structure/function relationship of YihS, we determined the crystal structure of S. enterica YihS mutant (H248A) in complex with a substrate (D-mannose) at 1.6 A resolution. This enzyme-substrate complex structure is the first demonstration in the AGE structural family, and it enables us to identify active-site residues and postulate a reaction mechanism for YihS. The substrate, beta-d-mannose, fits well in the active site and is specifically recognized by the enzyme. The substrate-binding site of YihS for the mannose C1 and O5 atoms is architecturally similar to those of mutarotases, suggesting that YihS adopts the pyranose ring-opening process by His383 and acidifies the C2 position, forming an aldehyde at the C1 position. In the isomerization step, His248 functions as a base catalyst responsible for transferring the proton from the C2 to C1 positions through a cis-enediol intermediate. On the other hand, in AGE, His248 is thought to abstract and re-adduct the proton at the C2 position of the substrate. These findings provide not only molecular insights into the YihS reaction mechanism but also useful information for the molecular design of novel carbohydrate-active enzymes with the common scaffold, alpha6/alpha6-barrel.
- Division of Food Science and Biotechnology, Graduate School of Agriculture, Kyoto University, Gokasho, Uji, Kyoto 611-0011, Japan.
Organizational Affiliation: