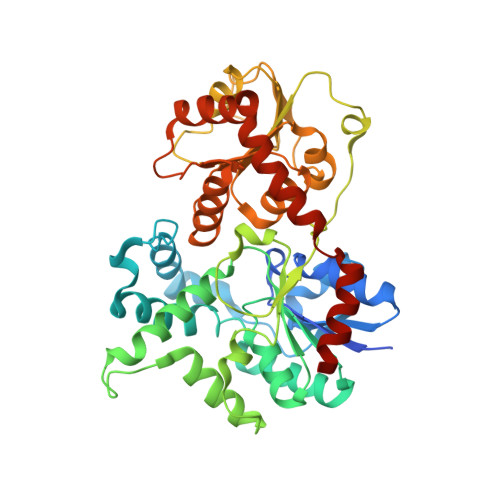
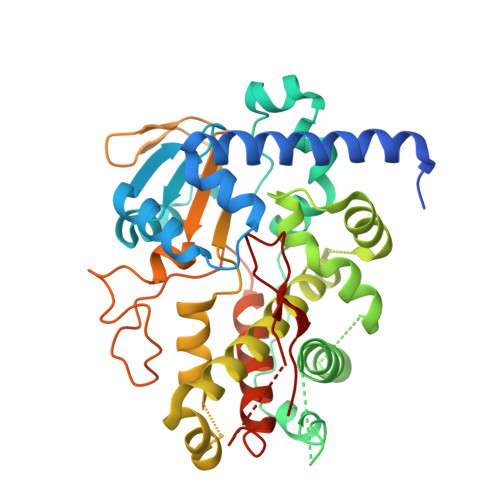
Structure of the Glycosyltransferase Eryciii in Complex with its Activating P450 Homologue Erycii.
Moncrieffe, M.C., Fernandez, M., Spiteller, D., Matsumura, H., Gay, N.J., Luisi, B.F., Leadlay, P.F.(2012) J Mol Biology 415: 92
- PubMed: 22056329 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2011.10.036
- Primary Citation Related Structures:
2YJN - PubMed Abstract:
In the biosynthesis of the clinically important antibiotic erythromycin D, the glycosyltransferase (GT) EryCIII, in concert with its partner EryCII, attaches a nucleotide-activated sugar to the macrolide scaffold with high specificity. To understand the role of EryCII, we have determined the crystal structure of the EryCIII·EryCII complex at 3.1 Å resolution. The structure reveals a heterotetramer with a distinctive, elongated quaternary organization. The EryCIII subunits form an extensive self-complementary dimer interface at the center of the complex, and the EryCII subunits lie on the periphery. EryCII binds in the vicinity of the putative macrolide binding site of EryCIII but does not make direct interactions with this site. Our biophysical and enzymatic data support a model in which EryCII stabilizes EryCIII and also functions as an allosteric activator of the GT.
- Department of Biochemistry, Cambridge University, Cambridge CB2 1GA, UK. mcm35@cam.ac.uk
Organizational Affiliation: