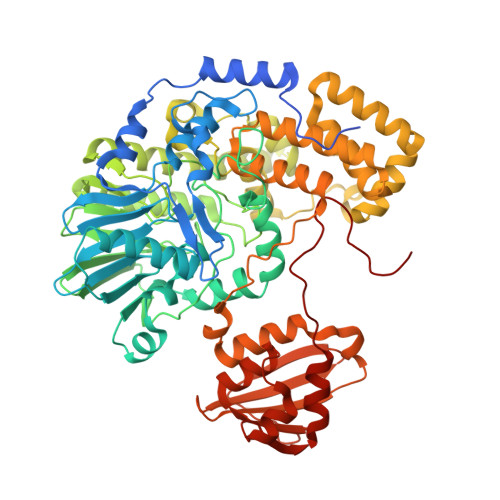
Structure and Mechanism of an Inverting Alkylsulfatase from Pseudomonas Sp. Dsm6611 Specific for Secondary Alkylsulfates.
Knaus, T., Schober, M., Kepplinger, B., Faccinelli, M., Pitzer, J., Faber, K., Macheroux, P., Wagner, U.(2012) FEBS J 279: 4374
- PubMed: 23061549
- DOI: https://doi.org/10.1111/febs.12027
- Primary Citation Related Structures:
2YHE, 4AV7, 4AXH - PubMed Abstract:
A highly enantioselective and stereoselective secondary alkylsulfatase from Pseudomonas sp. DSM6611 (Pisa1) was heterologously expressed in Escherichia coli BL21, and purified to homogeneity for kinetic and structural studies. Structure determination of Pisa1 by X-ray crystallography showed that the protein belongs to the family of metallo-β-lactamases with a conserved binuclear Zn(2+) cluster in the active site. In contrast to a closely related alkylsulfatase from Pseudomonas aeruginosa (SdsA1), Pisa1 showed a preference for secondary rather than primary alkyl sulfates, and enantioselectively hydrolyzed the (R)-enantiomer of rac-2-octyl sulfate, yielding (S)-2-octanol with inversion of absolute configuration as a result of C-O bond cleavage. In order to elucidate the mechanism of inverting sulfate ester hydrolysis, for which no counterpart in chemical catalysis exists, we designed variants of Pisa1 guided by three-dimensional structure and docking experiments. In the course of these studies, we identified an invariant histidine (His317) near the sulfate-binding site as the general acid for crucial protonation of the sulfate leaving group. Additionally, amino acid replacements in the alkyl chain-binding pocket generated an enzyme variant that lost its stereoselectivity towards rac-2-octyl sulfate. These findings are discussed in light of the potential use of this enzyme family for applications in biocatalysis.
- Institute of Biochemistry, Graz University of Technology, Graz, Austria.
Organizational Affiliation: