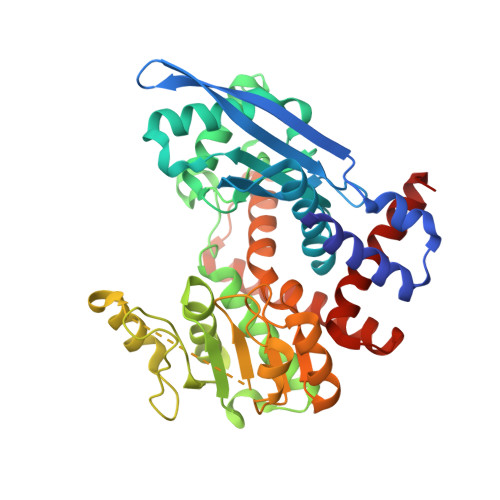
Crystal Structure of Nad-Dependent Peptoniphilus Asaccharolyticus Glutamate Dehydrogenase Reveals Determinants of Cofactor Specificity.
Oliveira, T., Panjikar, S., Carrigan, J.B., Hamza, M., Engel, P.C., Khan, A.R.(2012) J Struct Biol 177: 543
- PubMed: 22068154
- DOI: https://doi.org/10.1016/j.jsb.2011.10.006
- Primary Citation of Related Structures:
2YFQ - PubMed Abstract:
Glutamate dehydrogenases (EC 1.4.1.2-4) catalyse the oxidative deamination of l-glutamate to α-ketoglutarate using NAD(P) as a cofactor. The bacterial enzymes are hexamers and each polypeptide consists of an N-terminal substrate-binding (Domain I) followed by a C-terminal cofactor-binding segment (Domain II). The reaction takes place at the junction of the two domains, which move as rigid bodies and are presumed to narrow the cleft during catalysis. Distinct signature sequences in the nucleotide-binding domain have been linked to NAD(+) vs. NADP(+) specificity, but they are not unambiguous predictors of cofactor preferences. Here, we have determined the crystal structure of NAD(+)-specific Peptoniphilus asaccharolyticus glutamate dehydrogenase in the apo state. The poor quality of native crystals was resolved by derivatization with selenomethionine, and the structure was solved by single-wavelength anomalous diffraction methods. The structure reveals an open catalytic cleft in the absence of substrate and cofactor. Modeling of NAD(+) in Domain II suggests that a hydrophobic pocket and polar residues contribute to nucleotide specificity. Mutagenesis and isothermal titration calorimetry studies of a critical glutamate at the P7 position of the core fingerprint confirms its role in NAD(+) binding. Finally, the cofactor binding site is compared with bacterial and mammalian enzymes to understand how the amino acid sequences and three-dimensional structures may distinguish between NAD(+) vs. NADP(+) recognition.
- School of Biochemistry and Immunology, Trinity College Dublin, Dublin 2, Ireland.
Organizational Affiliation: