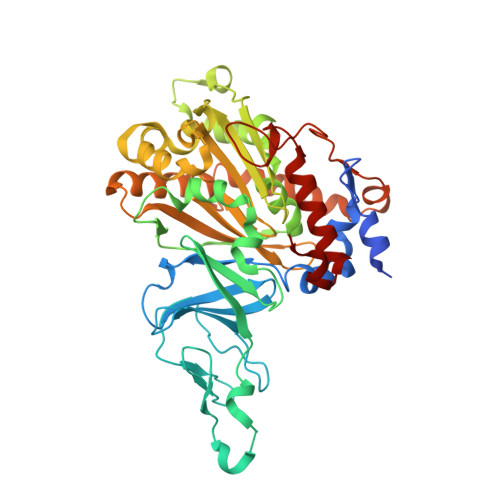
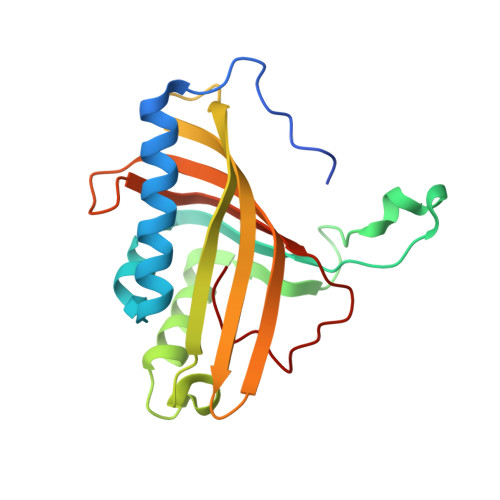
Retuning Rieske-Type Oxygenases to Expand Substrate Range.
Mohammadi, M., Viger, J., Kumar, P., Barriault, D., Bolin, J.T., Sylvestre, M.(2011) J Biological Chem 286: 27612
- PubMed: 21653696 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M111.255174
- Primary Citation Related Structures:
2YFI, 2YFJ - PubMed Abstract:
Rieske-type oxygenases are promising biocatalysts for the destruction of persistent pollutants or for the synthesis of fine chemicals. In this work, we explored pathways through which Rieske-type oxygenases evolve to expand their substrate range. BphAE(p4), a variant biphenyl dioxygenase generated from Burkholderia xenovorans LB400 BphAE(LB400) by the double substitution T335A/F336M, and BphAE(RR41), obtained by changing Asn(338), Ile(341), and Leu(409) of BphAE(p4) to Gln(338), Val(341), and Phe(409), metabolize dibenzofuran two and three times faster than BphAE(LB400), respectively. Steady-state kinetic measurements of single- and multiple-substitution mutants of BphAE(LB400) showed that the single T335A and the double N338Q/L409F substitutions contribute significantly to enhanced catalytic activity toward dibenzofuran. Analysis of crystal structures showed that the T335A substitution relieves constraints on a segment lining the catalytic cavity, allowing a significant displacement in response to dibenzofuran binding. The combined N338Q/L409F substitutions alter substrate-induced conformational changes of protein groups involved in subunit assembly and in the chemical steps of the reaction. This suggests a responsive induced fit mechanism that retunes the alignment of protein atoms involved in the chemical steps of the reaction. These enzymes can thus expand their substrate range through mutations that alter the constraints or plasticity of the catalytic cavity to accommodate new substrates or that alter the induced fit mechanism required to achieve proper alignment of reaction-critical atoms or groups.
- Institut National de la Recherche Scientifique-Institut Armand-Frappier, Laval, Quebec H7V 1B7, Canada.
Organizational Affiliation: