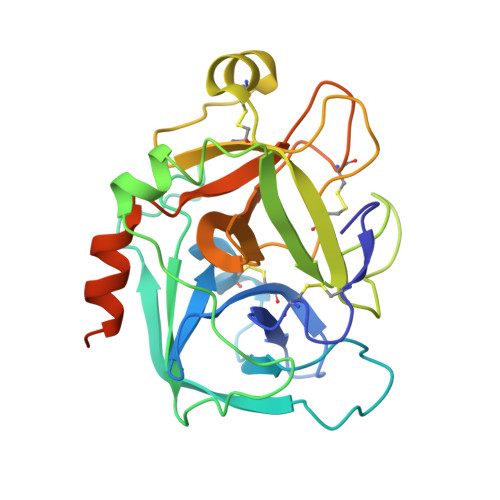
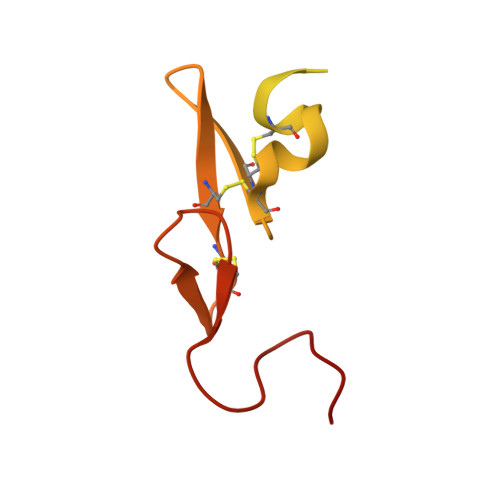
Structure and Property Based Design of Factor Xa Inhibitors: Pyrrolidin-2-Ones with Aminoindane and Phenylpyrrolidine P4 Motifs.
Young, R.J., Adams, C., Blows, M., Brown, D., Burns-Kurtis, C.L., Chan, C., Chaudry, L., Convery, M.A., Davies, D.E., Exall, A.M., Foster, G., Harling, J.D., Hortense, E., Irvine, S., Irving, W.R., Jackson, S., Kleanthous, S., Pateman, A.J., Patikis, A.N., Roethka, T.J., Senger, S., Stelman, G.J., Toomey, J.R., West, R.I., Whittaker, C., Zhou, P., Watson, N.S.(2011) Bioorg Med Chem Lett 21: 1582
- PubMed: 21349710 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2011.01.131
- Primary Citation Related Structures:
2Y7Z, 2Y80, 2Y81, 2Y82 - PubMed Abstract:
The rational design, syntheses and evaluation of potent sulfonamidopyrrolidin-2-one-based factor Xa inhibitors incorporating aminoindane and phenylpyrrolidine P4 motifs are described. These series delivered highly potent anticoagulant compounds with excellent oral pharmacokinetic profiles; however, significant time dependant P450 inhibition was an issue for the aminoindane series, but this was not observed with the phenylpyrrolidine motif, which produced candidate quality molecules with potential for once-daily oral dosing in humans.
- GlaxoSmithKline, Medicines Research Centre, Stevenage, Hertfordshire, United Kingdom. Rob.J.Young@gsk.com
Organizational Affiliation: