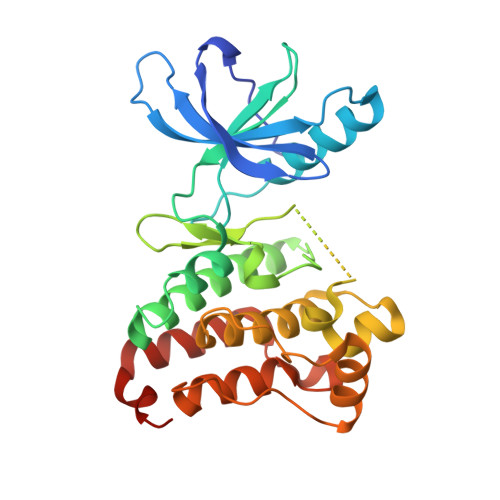
Crystal Structure of the Epha4 Protein Tyrosine Kinase Domain in the Apo- and Dasatinib-Bound State.
Farenc, C.J.A., Hameetman, L., Zoutman, W., Tensen, C.P., Siegal, G.(2011) FEBS Lett 585: 3593
- PubMed: 22036717 Search on PubMed
- DOI: https://doi.org/10.1016/j.febslet.2011.10.028
- Primary Citation Related Structures:
2Y6M, 2Y6O - PubMed Abstract:
The Eph family of receptor tyrosine kinases regulates diverse cellular processes while the over-expression of a member of this family, EphA4, has been reported in a variety of malignant carcinomas. To gain insight into molecular mechanisms and to facilitate structure-based inhibitor design, we solved the crystal structure of the native EphA4 kinase domain in both the apo and dasatinib bound forms. Analysis of the two structures provides insight into structural features of inhibitor binding and revealed a hydrophobic back-pocket in the ATP- binding site of EphA4 which was previously unidentified. The structures suggest a route towards development of novel and specific inhibitors.
- Protein Chemistry Group, Leiden Institute of Chemistry, Leiden University, Leiden, The Netherlands.
Organizational Affiliation: