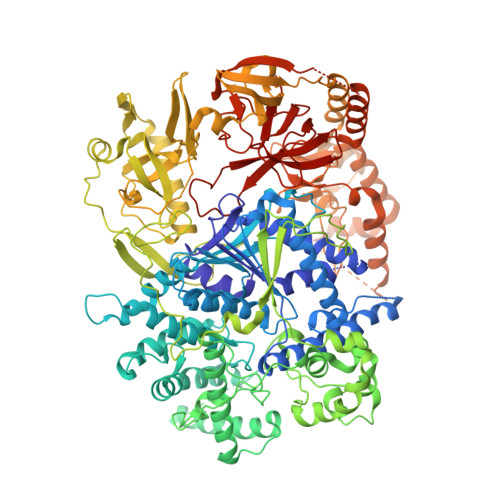
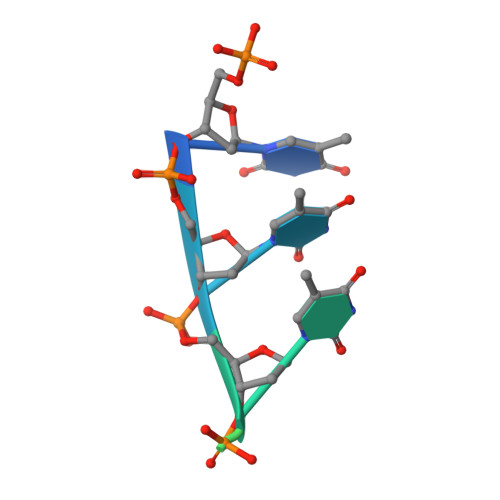
Coupled 5' Nucleotide Recognition and Processivity in Xrn1-Mediated Mrna Decay.
Jinek, M., Coyle, S.M., Doudna, J.A.(2011) Mol Cell 41: 600
- PubMed: 21362555 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.molcel.2011.02.004
- Primary Citation Related Structures:
2Y35 - PubMed Abstract:
Messenger RNA decay plays a central role in the regulation and surveillance of eukaryotic gene expression. The conserved multidomain exoribonuclease Xrn1 targets cytoplasmic RNA substrates marked by a 5' monophosphate for processive 5'-to-3' degradation by an unknown mechanism. Here, we report the crystal structure of an Xrn1-substrate complex. The single-stranded substrate is held in place by stacking of the 5'-terminal trinucleotide between aromatic side chains while a highly basic pocket specifically recognizes the 5' phosphate. Mutations of residues involved in binding the 5'-terminal nucleotide impair Xrn1 processivity. The substrate recognition mechanism allows Xrn1 to couple processive hydrolysis to duplex melting in RNA substrates with sufficiently long single-stranded 5' overhangs. The Xrn1-substrate complex structure thus rationalizes the exclusive specificity of Xrn1 for 5'-monophosphorylated substrates, ensuring fidelity of mRNA turnover, and posits a model for translocation-coupled unwinding of structured RNA substrates.
- Department of Molecular and Cell Biology, University of California, Berkeley, Berkeley, CA 94720, USA.
Organizational Affiliation: