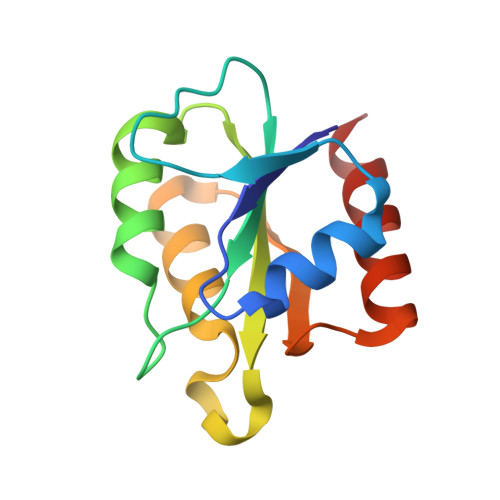
High Resolution Crystal Structures of Nrdi in the Oxidised and Reduced States: An Unusual Flavodoxin
Johansson, R., Torrents, E., Lundin, D., Sprenger, J., Sahlin, M., Sjoberg, B.M., Logan, D.T.(2010) FEBS J 277: 4265
- PubMed: 20831589
- DOI: https://doi.org/10.1111/j.1742-4658.2010.07815.x
- Primary Citation of Related Structures:
2XOD, 2XOE - PubMed Abstract:
The small flavoprotein NrdI is an essential component of the class Ib ribonucleotide reductase system in many bacteria. NrdI interacts with the class Ib radical generating protein NrdF. It is suggested to be involved in the rescue of inactivated diferric centres or generation of active dimanganese centres in NrdF. Although NrdI bears a superficial resemblance to flavodoxin, its redox properties have been demonstrated to be strikingly different. In particular, NrdI is capable of two-electron reduction, whereas flavodoxins are exclusively one-electron reductants. This has been suggested to depend on a lesser destabilization of the negatively-charged hydroquinone state than in flavodoxins. We have determined the crystal structures of NrdI from Bacillus anthracis, the causative agent of anthrax, in the oxidized and semiquinone forms, at resolutions of 0.96 and 1.4 Å, respectively. These structures, coupled with analysis of all curated NrdI sequences, suggest that NrdI defines a new structural family within the flavodoxin superfamily. The conformational behaviour of NrdI in response to FMN reduction is very similar to that of flavodoxins, involving a peptide flip in a loop near the N5 atom of the flavin ring. However, NrdI is much less negatively charged than flavodoxins, which is expected to affect its redox properties significantly. Indeed, sequence analysis shows a remarkable spread in the predicted isoelectric points of NrdIs, from approximately pH 4-10. The implications of these observations for class Ib ribonucleotide reductase function are discussed.
- Department of Biochemistry and Structural Biology, Lund University, Sweden.
Organizational Affiliation: