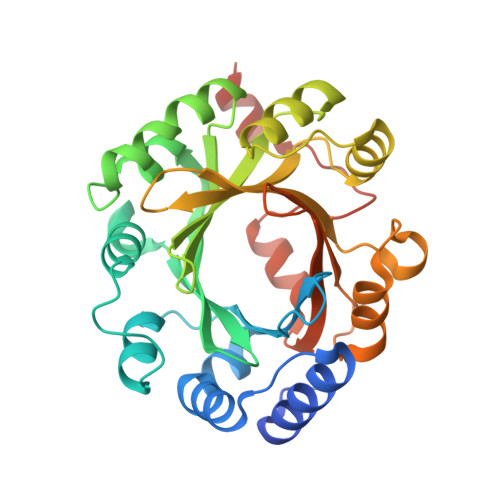
Structure and Mechanism of the Magnesium-Independent Aromatic Prenyltransferase Cloq from the Clorobiocin Biosynthetic Pathway.
Metzger, U., Keller, S., Stevenson, C.E.M., Heide, L., Lawson, D.M.(2010) J Mol Biology 404: 611
- PubMed: 20946900 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2010.09.067
- Primary Citation Related Structures:
2XLQ, 2XLY, 2XM5, 2XM7 - PubMed Abstract:
CloQ is an aromatic prenyltransferase from the clorobiocin biosynthetic pathway of Streptomyces roseochromogenes var. oscitans. It is involved in the synthesis of the prenylated hydroxybenzoate moiety of the antibiotic, specifically catalyzing the attachment of a dimethylallyl moiety to 4-hydroxyphenylpyruvate. Herein, we report the crystal structure of CloQ and use it as a framework for interpreting biochemical data from both wild-type and variant proteins. CloQ belongs to the aromatic prenyltransferase family, which is characterized by an unusual core fold comprising five consecutive ααββ elements that form a central 10-stranded anti-parallel β-barrel. The latter delineates a solvent-accessible cavity where substrates bind and catalysis takes place. This cavity has well-defined polar and nonpolar regions, which have distinct roles in substrate binding and facilitate a Friedel-Crafts-type mechanism. We propose that the juxtaposition of five positively charged residues in the polar region circumvents the necessity for a Mg(2+), which, by contrast, is a strict requirement for the majority of prenyltransferases characterized to date. Our structure of CloQ complexed with 4-hydroxyphenylpyruvate reveals the formation of a covalent link between the substrate and Cys215 to yield a thiohemiketal species. Through site-directed mutagenesis, we show that this link is not essential for enzyme activity in vitro. Furthermore, we demonstrate that CloQ will accept alternative substrates and, therefore, has the capacity to generate a range of prenylated compounds. Since prenylation is thought to enhance the bioactivity of many natural products, CloQ offers considerable promise as a biocatalyst for the chemoenzymatic synthesis of novel compounds with therapeutic potential.
- Department of Biological Chemistry, John Innes Centre, Norwich NR4 7UH, UK.
Organizational Affiliation: