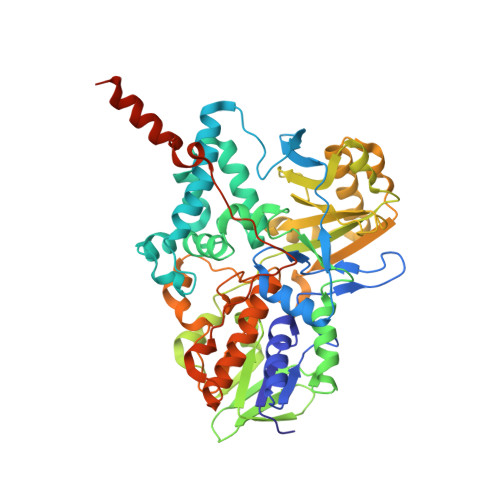
Potentiation of ligand binding through cooperative effects in monoamine oxidase B.
Bonivento, D., Milczek, E.M., McDonald, G.R., Binda, C., Holt, A., Edmondson, D.E., Mattevi, A.(2010) J Biological Chem 285: 36849-36856
- PubMed: 20855894 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M110.169482
- Primary Citation Related Structures:
2XCG, 2XFN, 2XFO, 2XFP, 2XFQ, 2XFU - PubMed Abstract:
Crystallographic and biochemical studies have been employed to identify the binding site and mechanism for potentiation of imidazoline binding in human monoamine oxidase B (MAO B). 2-(2-Benzofuranyl)-2-imidazoline (2-BFI) inhibits recombinant human MAO B with a K(i) of 8.3 ± 0.6 μM, whereas tranylcypromine-inhibited MAO B binds 2-BFI with a K(d) of 9 ± 2 nM, representing an increase in binding energy Δ(ΔG) of -3.9 kcal/mol. Crystal structures show the imidazoline ligand bound in a site that is distinct from the substrate-binding cavity. Contributions to account for the increase in binding affinity upon tranylcypromine inhibition include a conformational change in the side chain of Gln(206) and a "closed conformation" of the side chain of Ile(199), forming a hydrophobic "sandwich" with the side chain of Ile(316) on each face of the benzofuran ring of 2-BFI. Data with the I199A mutant of human MAO B and failure to observe a similar binding potentiation with rat MAO B, where Ile(316) is replaced with a Val residue, support an allosteric mechanism where the increased binding affinity of 2-BFI results from a cooperative increase in H-bond strength through formation of a more hydrophobic milieu. These insights should prove valuable in the design of high affinity and specific reversible MAO B inhibitors.
- Department of Genetics and Microbiology, University of Pavia, I-27100 Pavia PV, Italy.
Organizational Affiliation: