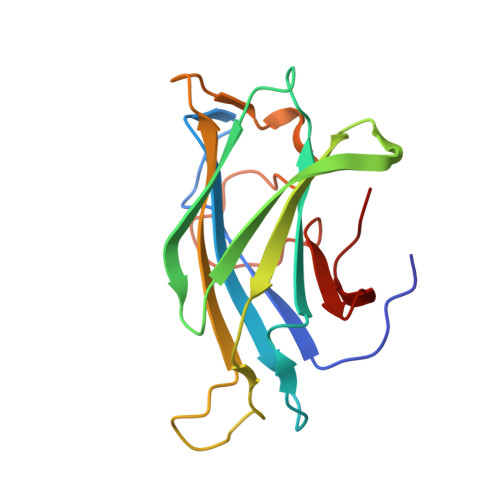
Scaffoldin-Borne Family 3B Carbohydrate-Binding Module from the Cellulosome of Bacteroides Cellulosolvens: Structural Diversity and Significance of Calcium for Carbohydrate Binding
Yaniv, O., Shimon, L.J.W., Bayer, E.A., Lamed, R., Frolow, F.(2011) Acta Crystallogr D Biol Crystallogr 67: 506
- PubMed: 21636890 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444911011322
- Primary Citation Related Structures:
2XBT - PubMed Abstract:
The potent cellulose-binding modules of cellulosomal scaffoldin subunits belong to the greater family of carbohydrate-binding modules (CBMs). They have generally been classified as belonging to family 3a on the basis of sequence similarity. They form nine-stranded β-sandwich structures with jelly-roll topology. The members of this family possess on their surface a planar array of aromatic amino-acid residues (known as the linear strip) that form stacking interactions with the glucose rings of cellulose chains and have a conserved Ca(2+)-binding site. Intriguingly, the CBM3 from scaffoldin A (ScaA) of Bacteroides cellulosolvens exhibits alterations in sequence that make it more similar to the CBMs of free cellulolytic enzymes, which are classified into CBM family 3b. X-ray structural analysis was undertaken in order to examine the structural consequences of the sequence changes and the consequent family affiliation. The CBM3 crystallized in space group I4(1)22 with one molecule in the asymmetric unit, yielding diffraction to a resolution of 1.83 Å using X-ray synchrotron radiation. Compared with the known structures of other scaffoldin-borne CBMs, a sequence insertion and deletion appear to compensate for each other as both contained an aromatic residue that is capable of contributing to cellulose binding; hence, even though there are alterations in the composition and localization of the aromatic residues in the linear strip its binding ability was not compromised. Interestingly, no Ca(2+) ions were detected in the conserved calcium-binding site, although the module was properly folded; this suggests that the structural role of Ca(2+) is less important than originally supposed. These observations indicate that despite their conserved function the scaffoldin-borne CBMs are more diverse in their sequences and structures than previously assumed.
- Department of Molecular Microbiology and Biotechnology, Tel Aviv University, Tel Aviv 69978, Israel.
Organizational Affiliation: