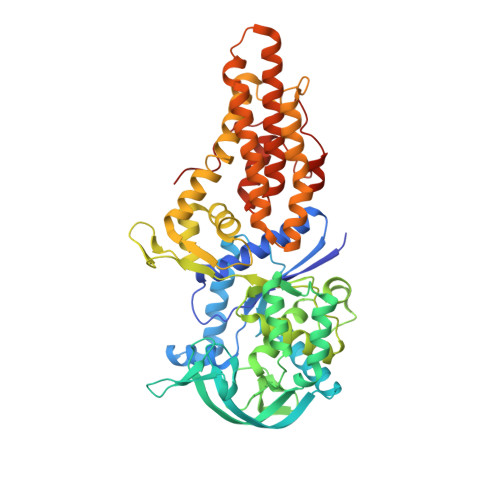
Flexibility and Communication within the Structure of the Mycobacterium Smegmatis Methionyl-tRNA Synthetase.
Ingvarsson, H., Unge, T.(2010) FEBS J 277: 3947
- PubMed: 20796028 Search on PubMed
- DOI: https://doi.org/10.1111/j.1742-4658.2010.07784.x
- Primary Citation Related Structures:
2X1L, 2X1M - PubMed Abstract:
Two structures of monomeric methionyl-tRNA synthetase, from Mycobacterium smegmatis, in complex with the ligands methionine/adenosine and methionine, were analyzed by X-ray crystallography at 2.3 Å and at 2.8 Å, respectively. The structures demonstrated the flexibility of the multidomain enzyme. A new conformation of the structure was identified in which the connective peptide domain bound more closely to the catalytic domain than described previously. The KMSKS(301-305) loop in our structures was in an open and inactive conformation that differed from previous structures by a rotation of the loop of about 90° around hinges located at Asn297 and Val310. The binding of adenosine to the methionyl-tRNA synthetase methionine complex caused a shift in the KMSKS domain that brought it closer to the catalytic domain. The potential use of the adenosine-binding site for inhibitor binding was evaluated and a potential binding site for a specific allosteric inhibitor was identified.
- Department of Cell and Molecular Biology, Uppsala Biomedical Center, Uppsala University, Uppsala, Sweden.
Organizational Affiliation: