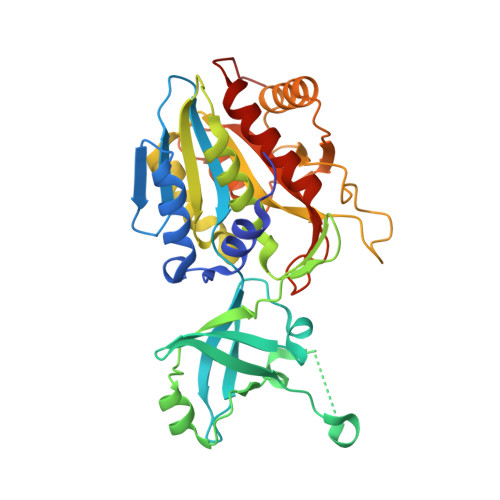
The Structural and Biochemical Characterizations of a Novel Tet Peptidase Complex from Pyrococcus Horikoshii Reveal an Integrated Peptide Degradation System in Hyperthermophilic Archaea.
Dura, M.A., Rosenbaum, E., Larabi, A., Gabel, F., Vellieux, F.M., Franzetti, B.(2009) Mol Microbiol 72: 26
- PubMed: 19291145
- DOI: https://doi.org/10.1111/j.1365-2958.2009.06600.x
- Primary Citation Related Structures:
2WZN - PubMed Abstract:
The structure of a 468 kDa peptidase complex from the hyperthermophile Pyrococcus horikoshii has been solved at 1.9 A resolution. The monomer contains the M42 peptidase typical catalytic domain, and a dimerization domain that allows the formation of dimers that assemble as a 12-subunit self-compartmentalized tetrahedron, similar to those described for the TET peptidases. The biochemical analysis shows that the enzyme is cobalt-activated and cleaves peptides by a non-processive mechanism. Consequently, this protein represents the third TET peptidase complex described in P. horikoshii, thereby called PhTET3. It is a lysyl aminopeptidase with a strong preference for basic residues, which are poorly cleaved by PhTET1 and PhTET2. The structural analysis of PhTET3 and its comparison with PhTET1 and PhTET2 unravels common features explaining the general mode of action of the TET molecular machines as well as differences that can be associated with strong substrate discriminations. The question of the stability of the TET assemblies under extreme temperatures has been addressed. PhTET3 displays its maximal activity at 95 degrees C and small-angle neutron scattering experiments at 90 degrees C demonstrate the absence of quaternary structure alterations after extensive incubation times. In conclusion, PhTETs are complementary peptide destruction machines that may play an important role in the metabolism of P. horikoshii.
- Institut de Biologie Structurale J.-P. Ebel, UMR 5075 CNRS-CEA-UJF, 41 rue Jules Horowitz, 38027 Grenoble, France.
Organizational Affiliation: