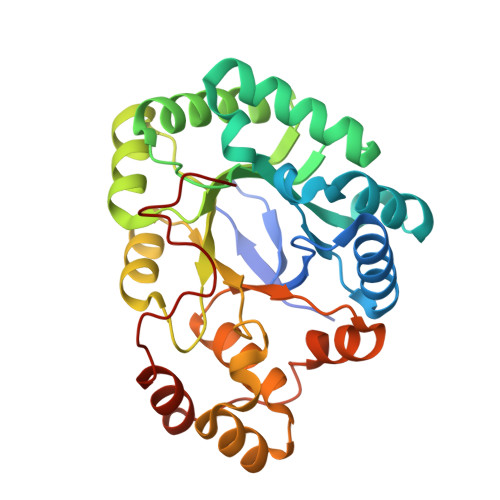
Crystal Structure and Comparative Functional Analyses of a Mycobacterium Aldo-Keto Reductase.
Scoble, J., Mcalister, A.D., Fulton, Z., Troy, S., Byres, E., Vivian, J.P., Brammananth, R., Wilce, M.C.J., Le Nours, J., Zaker-Tabrizi, L., Coppel, R.L., Crellin, P.K., Rossjohn, J., Beddoe, T.(2010) J Mol Biology 398: 26
- PubMed: 20188740
- DOI: https://doi.org/10.1016/j.jmb.2010.02.021
- Primary Citation of Related Structures:
2WZM, 2WZT - PubMed Abstract:
Aldo-keto reductases (AKRs) are a large superfamily of NADPH-dependent enzymes that catalyze the reduction of aldehydes, aldoses, dicarbonyls, steroids, and monosaccharides. While their precise physiological role is generally unknown, AKRs are nevertheless involved in the detoxification of a broad range of toxic metabolites. Mycobacteria contain a number of AKRs, the majority of which are uncharacterised. Here, we report the 1.9 and 1.6 A resolution structures of the apoenzyme and NADPH-bound forms, respectively, of an AKR (MSMEG_2407) from Mycobacterium smegmatis, a close homologue of the M. tuberculosis enzyme Rv2971, whose function is essential to this bacterium. MSMEG_2407 adopted the triosephosphate isomerase (alpha/beta)(8)-barrel fold exhibited by other AKRs. MSMEG_2407 (AKR5H1) bound NADPH via an induced-fit mechanism, in which the NADPH was ligated in an extended fashion. Polar-mediated interactions dominated the interactions with the cofactor, which is atypical of the mode of NADPH binding within the AKR family. Moreover, the nicotinamide ring of NADPH was disordered, and this was attributed to the lack of an "AKR-conserved" bulky residue within the nicotinamide-binding cavity of MSMEG_2407. Enzymatic characterisation of MSMEG_2407 and Rv2971 identified dicarbonyls as a preferred substrate family for hydrolysis, and the frontline antituberculosis drug isoniazid (INH) was shown to inhibit the enzyme activity of both recombinant MSMEG_2407 and Rv2971. However, differences between the affinities of MSMEG_2407 and Rv2971 for dicarbonyls and INH were observed, and this was attributable to amino acid substitutions within the cofactor- and substrate-binding sites. The structures of MSMEG_2407 and the accompanying biochemical characterisation of MSMEG_2407 and Rv2971 provide insight into the structure and function of AKRs from mycobacteria.
- Protein Crystallography Unit, Department of Biochemistry and Molecular Biology, School of Biomedical Sciences, Monash University, Clayton, Victoria 3800, Australia.
Organizational Affiliation: