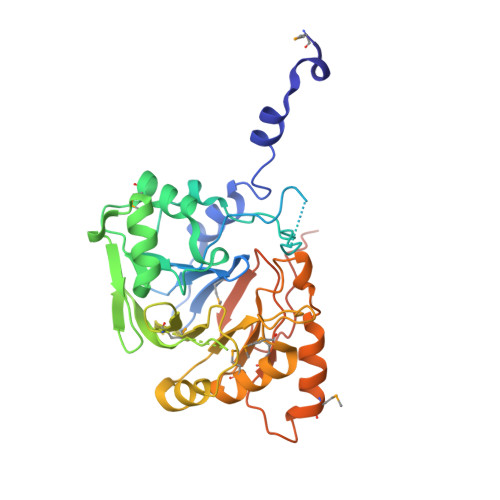
Molecular Architecture of the Mn(2+)Dependent Lactonase Ulag Reveals an Rnase-Like Metallo-Beta-Lactamase Fold and a Novel Quaternary Structure.
Garces, F., Fernandez, F.J., Montella, C., Penya-Soler, E., Prohens, R., Aguilar, J., Baldoma, L., Coll, M., Badia, J., Vega, M.C.(2010) J Mol Biology 398: 715
- PubMed: 20359483 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2010.03.041
- Primary Citation Related Structures:
2WYL, 2WYM - PubMed Abstract:
The ulaG gene, located in the ula regulon, is crucial for the catabolism of l-ascorbate under anaerobic conditions and it has been proposed to encode for the putative l-ascorbate-6-P lactonase. The ulaG gene is widespread among eubacteria, including human commensal and pathogenic genera such as Escherichia, Shigella, Klebsiella and Salmonella. Here, we report the three-dimensional structures of the apoenzyme and Mn(2+) holoenzyme of UlaG from E. coli to 2.6 A resolution, determined using single-wavelength anomalous diffraction phasing and molecular replacement, respectively. The structures reveal a highly specialized metallo-beta-lactamase-like fold derived from an ancient structural template that was involved in RNA maturation and DNA repair. This fold has a novel quaternary architecture consisting of a hexameric ring formed by a trimer of UlaG dimers. A mononuclear Mn(2)(+)-binding site resides at the core of the active site, which displays micromolar affinity for Mn(2+) and a distorted trigonal bipyramidal coordination. The active site Mn(2+) ion can be replaced by Co(2+) or Zn(2+), but not by Fe(3+). We further show that the Mn(2+) or Co(2)(+)-loaded enzyme exhibits lactonase activity towards l-ascorbate 6-P, thereby providing the first direct evidence of its catalytic role in the L-ascorbate catabolic pathway. Guided by the structural homology, we show that UlaG is able to cleave phosphodiester linkages in cyclic nucleotides, suggesting that the conservation of the fold and of the key catalytic residues allows for the evolutionary acquisition of substrate specificity for novel but related substrates.
- Department of Biochemistry and Molecular Biology, Institut de Biomedicina de la Universitat de Barcelona (IBUB), Faculty of Pharmacy, University of Barcelona, Barcelona, ES 08028, Spain.
Organizational Affiliation: