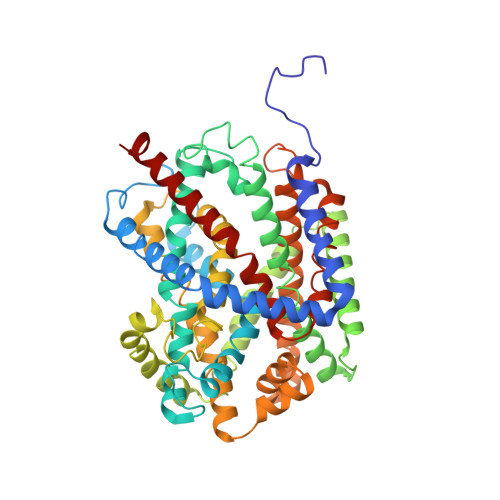
Structural Basis of Na(+)-Independent and Cooperative Substrate/Product Antiport in Cait.
Schulze, S., Koster, S., Geldmacher, U., Terwisscha Van Scheltinga, A.C., Kuhlbrandt, W.(2010) Nature 467: 233
- PubMed: 20829798 Search on PubMed
- DOI: https://doi.org/10.1038/nature09310
- Primary Citation Related Structures:
2WSW, 2WSX - PubMed Abstract:
Transport of solutes across biological membranes is performed by specialized secondary transport proteins in the lipid bilayer, and is essential for life. Here we report the structures of the sodium-independent carnitine/butyrobetaine antiporter CaiT from Proteus mirabilis (PmCaiT) at 2.3-A and from Escherichia coli (EcCaiT) at 3.5-A resolution. CaiT belongs to the family of betaine/carnitine/choline transporters (BCCT), which are mostly Na(+) or H(+) dependent, whereas EcCaiT is Na(+) and H(+) independent. The three-dimensional architecture of CaiT resembles that of the Na(+)-dependent transporters LeuT and BetP, but in CaiT a methionine sulphur takes the place of the Na(+) ion to coordinate the substrate in the central transport site, accounting for Na(+)-independent transport. Both CaiT structures show the fully open, inward-facing conformation, and thus complete the set of functional states that describe the alternating access mechanism. EcCaiT contains two bound butyrobetaine substrate molecules, one in the central transport site, the other in an extracellular binding pocket. In the structure of PmCaiT, a tryptophan side chain occupies the transport site, and access to the extracellular site is blocked. Binding of both substrates to CaiT reconstituted into proteoliposomes is cooperative, with Hill coefficients up to 1.7, indicating that the extracellular site is regulatory. We propose a mechanism whereby the occupied regulatory site increases the binding affinity of the transport site and initiates substrate translocation.
- Department of Structural Biology, Max Planck Institute of Biophysics, Max-von-Laue Strasse 3, 60438 Frankfurt am Main, Germany.
Organizational Affiliation: