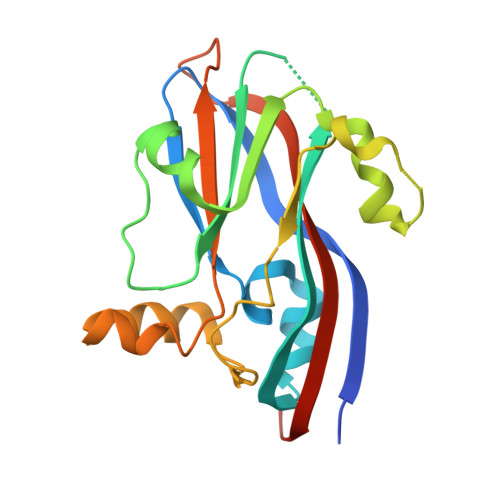
A Single Point Mutation Disrupts the Capsid Assembly in Sesbania Mosaic Virus Resulting in a Stable Isolated Dimer.
Pappachan, A., Chinnathambi, S., Satheshkumar, P.S., Savithri, H.S., Murthy, M.R.N.(2009) Virology 392: 215
- PubMed: 19643453 Search on PubMed
- DOI: https://doi.org/10.1016/j.virol.2009.06.047
- Primary Citation Related Structures:
2WLP - PubMed Abstract:
Protein-protein interactions play a crucial role in virus assembly and stability. With the view of disrupting capsid assembly and capturing smaller oligomers, interfacial residue mutations were carried out in the coat protein gene of Sesbania Mosaic Virus, a T=3 ss (+) RNA plant virus. A single point mutation of a Trp 170 present at the five-fold interface of the virus to a charged residue (Glu or Lys) arrested assembly of virus like particles and resulted in stable soluble dimers of the capsid protein. The X-ray crystal structure of one of the isolated dimer mutants - rCPDeltaN65W170K was determined to a resolution of 2.65 A. Detailed analysis of the dimeric mutant protein structure revealed that a number of structural changes take place, especially in the loop and interfacial regions during the course of assembly. The isolated dimer was "more relaxed" than the dimer found in the T=3 or T=1 capsids. The isolated dimer does not bind Ca(2+) ion and consequently four C-terminal residues are disordered. The FG loop, which interacts with RNA in the virus, has different conformations in the isolated dimer and the intact virus suggesting its flexible nature and the conformational changes that accompany assembly. The isolated dimer mutant was much less stable when compared to the assembled capsids, suggesting the importance of inter-subunit interactions and Ca(2+) mediated interactions in the stability of the capsids. With this study, SeMV becomes the first icosahedral virus for which X-ray crystal structures of T=3, T=1 capsids as well as a smaller oligomer of the capsid protein have been determined.
- Molecular Biophysics Unit, Indian Institute of Science, Bangalore, India.
Organizational Affiliation: