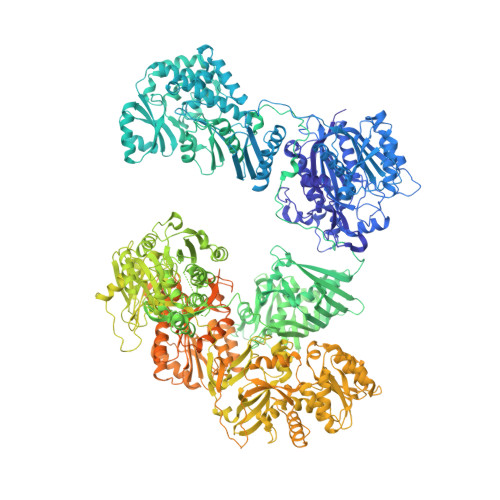
The Crystal Structure of a Mammalian Fatty Acid Synthase.
Maier, T., Leibundgut, M., Ban, N.(2008) Science 321: 1315
- PubMed: 18772430 Search on PubMed
- DOI: https://doi.org/10.1126/science.1161269
- Primary Citation Related Structures:
2VZ8, 2VZ9 - PubMed Abstract:
Mammalian fatty acid synthase is a large multienzyme that catalyzes all steps of fatty acid synthesis. We have determined its crystal structure at 3.2 angstrom resolution covering five catalytic domains, whereas the flexibly tethered terminal acyl carrier protein and thioesterase domains remain unresolved. The structure reveals a complex architecture of alternating linkers and enzymatic domains. Substrate shuttling is facilitated by flexible tethering of the acyl carrier protein domain and by the limited contact between the condensing and modifying portions of the multienzyme, which are mainly connected by linkers rather than direct interaction. The structure identifies two additional nonenzymatic domains: (i) a pseudo-ketoreductase and (ii) a peripheral pseudo-methyltransferase that is probably a remnant of an ancestral methyltransferase domain maintained in some related polyketide synthases. The structural comparison of mammalian fatty acid synthase with modular polyketide synthases shows how their segmental construction allows the variation of domain composition to achieve diverse product synthesis.
- Institute of Molecular Biology and Biophysics, ETH Zurich, 8092 Zurich, Switzerland.
Organizational Affiliation: