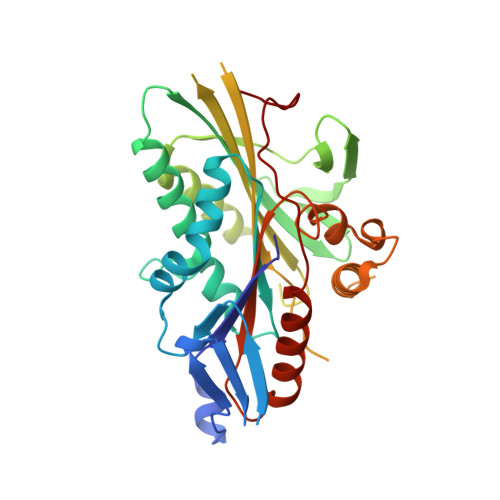
High-Resolution Crystal Structure and in Vivo Function of a Kinesin-2 Homologue in Giardia Intestinalis.
Hoeng, J.C., Dawson, S.C., House, S.A., Sagolla, M.S., Pham, J.K., Mancuso, J.J., Loewe, J., Cande, W.Z.(2008) Mol Biol Cell 19: 3124
- PubMed: 18463165
- DOI: https://doi.org/10.1091/mbc.e07-11-1156
- Primary Citation Related Structures:
2VVG - PubMed Abstract:
A critical component of flagellar assembly, the kinesin-2 heterotrimeric complex powers the anterograde movement of proteinaceous rafts along the outer doublet of axonemes in intraflagellar transport (IFT). We present the first high-resolution structures of a kinesin-2 motor domain and an ATP hydrolysis-deficient motor domain mutant from the parasitic protist Giardia intestinalis. The high-resolution crystal structures of G. intestinalis wild-type kinesin-2 (GiKIN2a) motor domain, with its docked neck linker and the hydrolysis-deficient mutant GiKIN2aT104N were solved in a complex with ADP and Mg(2+) at 1.6 and 1.8 A resolutions, respectively. These high-resolution structures provide unique insight into the nucleotide coordination within the active site. G. intestinalis has eight flagella, and we demonstrate that both kinesin-2 homologues and IFT proteins localize to both cytoplasmic and membrane-bound regions of axonemes, with foci at cell body exit points and the distal flagellar tips. We demonstrate that the T104N mutation causes GiKIN2a to act as a rigor mutant in vitro. Overexpression of GiKIN2aT104N results in significant inhibition of flagellar assembly in the caudal, ventral, and posterolateral flagellar pairs. Thus we confirm the conserved evolutionary structure and functional role of kinesin-2 as the anterograde IFT motor in G. intestinalis.
- Medical Research Council Laboratory of Molecular Biology, Cambridge CB2 2QH, United Kingdom.
Organizational Affiliation: