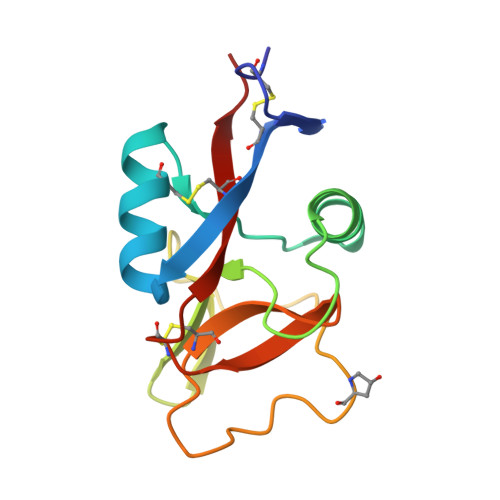
High Affinity Interaction between a Bivalve C-Type Lectin and a Biantennary Complex-Type N-Glycan Revealed by Crystallography and Microcalorimetry.
Gourdine, J.P., Cioci, G.C., Miguet, L., Unverzagt, C., Silva, D.V., Varrot, A., Gautier, C., Smith-Ravin, E.J., Imberty, A.(2008) J Biological Chem 283: 30112
- PubMed: 18687680
- DOI: https://doi.org/10.1074/jbc.M804353200
- Primary Citation of Related Structures:
2VUV, 2VUZ - PubMed Abstract:
Codakine is an abundant 14-kDa mannose-binding C-type lectin isolated from the gills of the sea bivalve Codakia orbicularis. Binding studies using inhibition of hemagglutination indicated specificity for mannose and fucose monosaccharides. Further experiments using a glycan array demonstrated, however, a very fine specificity for N-linked biantennary complex-type glycans. An unusually high affinity was measured by titration microcalorimetry performed with a biantennary Asn-linked nonasaccharide. The crystal structure of the native lectin at 1.3A resolution revealed a new type of disulfide-bridged homodimer. Each monomer displays three intramolecular disulfide bridges and contains only one calcium ion located in the canonical binding site that is occupied by a glycerol molecule. The structure of the complex between Asn-linked nonasaccharide and codakine has been solved at 1.7A resolution. All residues could be located in the electron density map, except for the capping beta1-4-linked galactosides. The alpha1-6-linked mannose binds to calcium by coordinating the O3 and O4 hydroxyl groups. The GlcNAc moiety of the alpha1,6 arm engages in several hydrogen bonds with the protein, whereas the GlcNAc on the other antenna is stacked against Trp(108), forming an extended binding site. This is the first structural report for a bivalve lectin.
- Département de Biologie, Université des Antilles et de la Guyane, Pointe-à-Pitre, F-97159 Guadeloupe, France.
Organizational Affiliation: