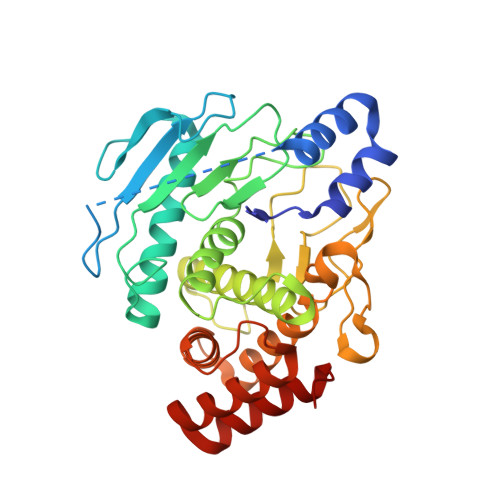
Crystal Structure of the Protealysin Precursor: Insights Into Propeptide Function.
Demidyuk, I.V., Gromova, T.Y., Polyakov, K.M., Melik-Adamyan, W.R., Kuranova, I.P., Kostrov, S.V.(2010) J Biological Chem 285: 2003
- PubMed: 19915005
- DOI: https://doi.org/10.1074/jbc.M109.015396
- Primary Citation Related Structures:
2VQX - PubMed Abstract:
Protealysin (PLN) belongs to the M4 family of peptidases that are commonly known as thermolysin-like proteases (TLPs). All TLPs are synthesized as precursors containing N-terminal propeptides. According to the primary structure of the N-terminal propeptides, the family is divided into two distinct groups. Representatives of the first group including thermolysin and all TLPs with known three-dimensional structures have long prosequences ( approximately 200 amino acids). Enzymes of the second group, whose prototype is protealysin, have short ( approximately 50 amino acids) propeptides. Here, we present the 1.8 A crystal structure of PLN precursor (proPLN), which is the first three-dimensional structure of a TLP precursor. Whereas the structure of the catalytic domain of proPLN is similar overall to previously reported structures of mature TLPs, it has specific features, including the absence of calcium-binding sites, and different structures of the N-terminal region and substrate-binding site. PLN propeptide forms a separate domain in the precursor and likely acts as an inhibitor that blocks the substrate-binding site and fixes the "open" conformation of the active site, which is unfavorable for catalysis. Furthermore the conserved PPL motif identified in our previous studies directly interacts with the S' subsites of the active center being a critical element of the propeptide-catalytic domain interface. Comparison of the primary structures of TLPs with short propeptides suggests that the specific features revealed in the proPLN crystal structure are typical for all protealysin-like enzymes. Thus, such proteins can be considered as a separate subfamily of TLPs.
- Institute of Molecular Genetics, Russian Academy of Sciences, Kurchatov Sq. 2, Moscow 123182, Russia. duk@img.ras.ru
Organizational Affiliation: