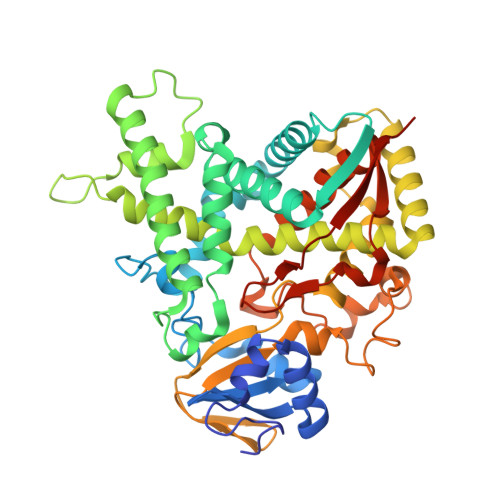
X-Ray Structure of 4,4'-Dihydroxybenzophenone Mimicking Sterol Substrate in the Active Site of Sterol 14Alpha-Demethylase (Cyp51)
Eddine, A.N., P Von Kries, J., Podust, M.V., Warrier, T., Kaufmann, S.H.E., Podust, L.M.(2008) J Biological Chem 283: 15152
- PubMed: 18367444 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M801145200
- Primary Citation Related Structures:
2VKU - PubMed Abstract:
A universal step in the biosynthesis of membrane sterols and steroid hormones is the oxidative removal of the 14alpha-methyl group from sterol precursors by sterol 14alpha-demethylase (CYP51). This enzyme is a primary target in treatment of fungal infections in organisms ranging from humans to plants, and development of more potent and selective CYP51 inhibitors is an important biological objective. Our continuing interest in structural aspects of substrate and inhibitor recognition in CYP51 led us to determine (to a resolution of 1.95A) the structure of CYP51 from Mycobacterium tuberculosis (CYP51(Mt)) co-crystallized with 4,4'-dihydroxybenzophenone (DHBP), a small organic molecule previously identified among top type I binding hits in a library screened against CYP51(Mt). The newly determined CYP51(Mt)-DHBP structure is the most complete to date and is an improved template for three-dimensional modeling of CYP51 enzymes from fungal and prokaryotic pathogens. The structure demonstrates the induction of conformational fit of the flexible protein regions and the interactions of conserved Phe-89 essential for both fungal drug resistance and catalytic function, which were obscure in the previously characterized CYP51(Mt)-estriol complex. DHBP represents a benzophenone scaffold binding in the CYP51 active site via a type I mechanism, suggesting (i) a possible new class of CYP51 inhibitors targeting flexible regions, (ii) an alternative catalytic function for bacterial CYP51 enzymes, and (iii) a potential for hydroxybenzophenones, widely distributed in the environment, to interfere with sterol biosynthesis. Finally, we show the inhibition of M. tuberculosis growth by DHBP in a mouse macrophage model.
- Max-Planck-Institute for Infection Biology, Berlin, Germany.
Organizational Affiliation: