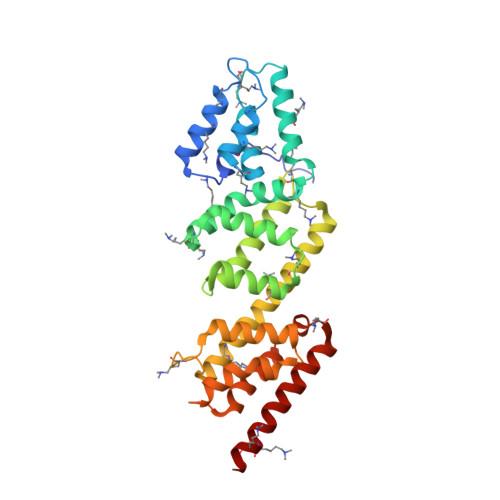
Structures of the Shigella Flexneri Type 3 Secretion System Protein Mxic Reveal Conformational Variability Amongst Homologues.
Deane, J.E., Roversi, P., King, C., Johnson, S., Lea, S.M.(2008) J Mol Biology 377: 985
- PubMed: 18304577
- DOI: https://doi.org/10.1016/j.jmb.2008.01.072
- Primary Citation of Related Structures:
2VIX, 2VJ4, 2VJ5 - PubMed Abstract:
Many Gram-negative pathogenic bacteria use a complex macromolecular machine, known as the type 3 secretion system (T3SS), to transfer virulence proteins into host cells. The T3SS is composed of a cytoplasmic bulb, a basal body spanning the inner and outer bacterial membranes, and an extracellular needle. Secretion is regulated by both cytoplasmic and inner membrane proteins that must respond to specific signals in order to ensure that virulence proteins are not secreted before contact with a eukaryotic cell. This negative regulation is mediated, in part, by a family of proteins that are thought to physically block the entrance to the secretion apparatus until an appropriate signal is received following host cell contact. Despite weak sequence homology between proteins of this family, the crystal structures of Shigella flexneri MxiC we present here confirm the conservation of domain topology with the homologue from Yersinia sp. Interestingly, comparison of the Shigella and Yersinia structures reveals a significant structural change that results in substantial domain re-arrangement and opening of one face of the molecule. The conservation of a negatively charged patch on this face suggests it may have a role in binding other components of the T3SS.
- Sir William Dunn School of Pathology, South Parks Rd, University of Oxford, Oxford OX1 3RE, UK.
Organizational Affiliation: