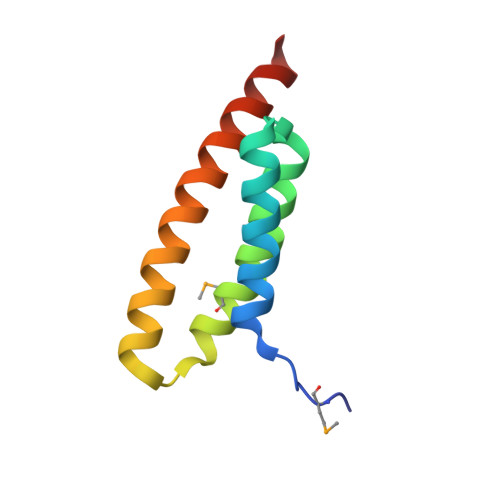
Structural Basis for Selective Recognition of Escrt-III by the Aaa ATPase Vps4
Obita, T., Saksena, S., Ghazi-Tabatabai, S., Gill, D.J., Perisic, O., Emr, S.D., Williams, R.L.(2007) Nature 449: 735
- PubMed: 17928861
- DOI: https://doi.org/10.1038/nature06171
- Primary Citation of Related Structures:
2V6X, 2V6Y - PubMed Abstract:
The AAA+ ATPases are essential for various activities such as membrane trafficking, organelle biogenesis, DNA replication, intracellular locomotion, cytoskeletal remodelling, protein folding and proteolysis. The AAA ATPase Vps4, which is central to endosomal traffic to lysosomes, retroviral budding and cytokinesis, dissociates ESCRT complexes (the endosomal sorting complexes required for transport) from membranes. Here we show that, of the six ESCRT--related subunits in yeast, only Vps2 and Did2 bind the MIT (microtubule interacting and transport) domain of Vps4, and that the carboxy-terminal 30 residues of the subunits are both necessary and sufficient for interaction. We determined the crystal structure of the Vps2 C terminus in a complex with the Vps4 MIT domain, explaining the basis for selective ESCRT-III recognition. MIT helices alpha2 and alpha3 recognize a (D/E)xxLxxRLxxL(K/R) motif, and mutations within this motif cause sorting defects in yeast. Our crystal structure of the amino-terminal domain of an archaeal AAA ATPase of unknown function shows that it is closely related to the MIT domain of Vps4. The archaeal ATPase interacts with an archaeal ESCRT-III-like protein even though these organisms have no endomembrane system, suggesting that the Vps4/ESCRT-III partnership is a relic of a function that pre-dates the divergence of eukaryotes and Archaea.
- MRC Laboratory of Molecular Biology, Medical Research Council Centre, Cambridge CB2 0QH, UK.
Organizational Affiliation: