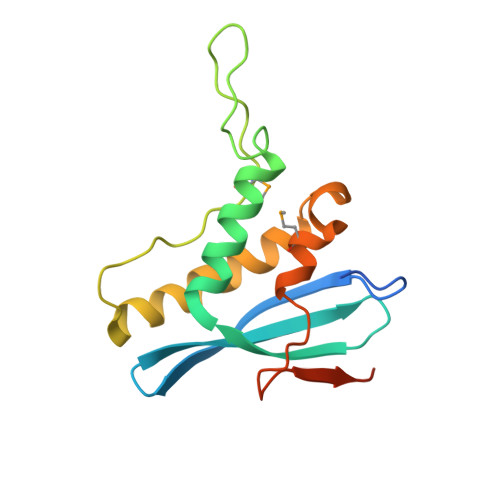
Structural and Membrane Binding Analysis of the Phox Homology Domain of Bem1P: Basis of Phosphatidylinositol 4-Phosphate Specificity.
Stahelin, R.V., Karathanassis, D., Murray, D., Williams, R.L., Cho, W.(2007) J Biological Chem 282: 25737
- PubMed: 17581820
- DOI: https://doi.org/10.1074/jbc.M702861200
- Primary Citation Related Structures:
2V6V - PubMed Abstract:
Phox homology (PX) domains, which have been identified in a variety of proteins involved in cell signaling and membrane trafficking, have been shown to interact with phosphoinositides (PIs) with different affinities and specificities. To elucidate the structural origin of the diverse PI specificity of PX domains, we determined the crystal structure of the PX domain from Bem1p that has been reported to bind phosphatidylinositol 4-phosphate (PtdIns(4)P). We also measured the membrane binding properties of the PX domain and its mutants by surface plasmon resonance and monolayer techniques and calculated the electrostatic potentials for the PX domain in the absence and presence of bound PtdIns(4)P. The Bem1p PX domain contains a signature PI-binding site optimized for PtdIns(4)P binding and also harbors basic and hydrophobic residues on the membrane-binding surface. The membrane binding of the Bem1p PX domain is initiated by nonspecific electrostatic interactions between the cationic membrane-binding surface of the domain and anionic membrane surfaces, followed by the membrane penetration of hydrophobic residues. Unlike other PX domains, the Bem1p PX domain has high intrinsic membrane penetrating activity in the absence of PtdIns(4)P, suggesting that the partial membrane penetration may occur before specific PtdIns(4)P binding and last after the removal of PtdIns(4)P under certain conditions. This structural and functional study of the PtdIns(4)P-binding Bem1p PX domain provides new insight into the diverse PI specificities and membrane-binding mechanisms of PX domains.
- Department of Chemistry, University of Illinois, Chicago, Illinois 60607-7061, USA.
Organizational Affiliation: