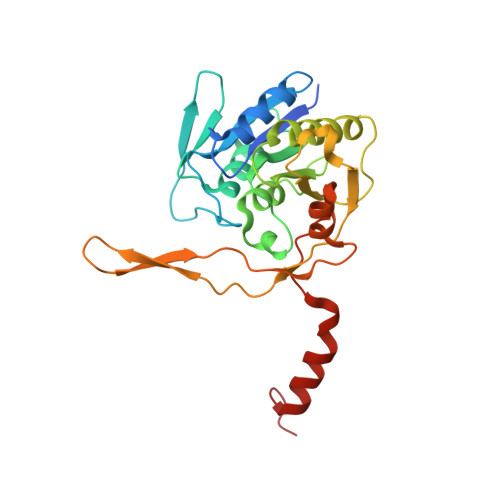
Structural and Functional Studies on a Mesophilic Stationary Phase Survival Protein (Sur E) from Salmonella Typhimurium
Pappachan, A., Savithri, H.S., Murthy, M.R.N.(2008) FEBS J 275: 5855
- PubMed: 19021761
- DOI: https://doi.org/10.1111/j.1742-4658.2008.06715.x
- Primary Citation Related Structures:
2V4N, 2V4O - PubMed Abstract:
SurE, the stationary-phase survival protein of Salmonella typhimurium, forms part of a stress survival operon regulated by the stationary-phase RNA polymerase alternative sigma factor. SurE is known to improve bacterial viability during stress conditions. It functions as a phosphatase specific to nucleoside monophosphates. In the present study we reported the X-ray crystal structure of SurE from Salmonella typhimurium. The protein crystallized in two forms: orthorhombic F222; and monoclinic C2. The two structures were determined to resolutions of 1.7 and 2.7 A, respectively. The protein exists as a domain-swapped dimer. The residue D230 is involved in several interactions that are probably crucial for domain swapping. A divalent metal ion is found at the active site of the enzyme, which is consistent with the divalent metal ion-dependent activity of the enzyme. Interactions of the conserved DD motif present at the N-terminus with the phosphate and the Mg(2+) present in the active site suggest that these residues play an important role in enzyme activity. The divalent metal ion specificity and the kinetic constants of SurE were determined using the generic phosphatase substrate para-nitrophenyl phosphate. The enzyme was inactive in the absence of divalent cations and was most active in the presence of Mg(2+). Thermal denaturation studies showed that S. typhimurium SurE is much less stable than its homologues and an attempt was made to understand the molecular basis of the lower thermal stability based on solvation free-energy. This is the first detailed crystal structure analysis of SurE from a mesophilic organism.
- Molecular Biophysics Unit, Indian Institute of Science, Bangalore, India.
Organizational Affiliation: