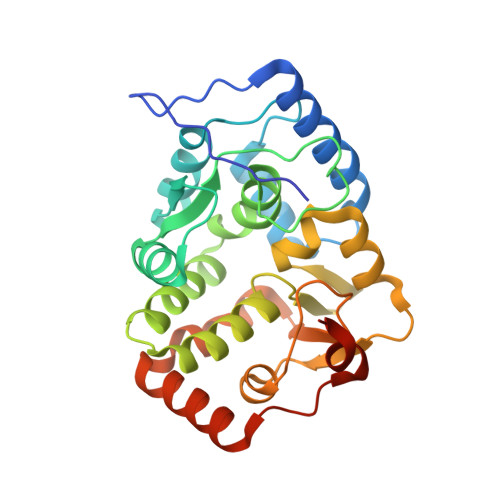
Structure of Phenylalanine Hydroxylase from Colwellia Psychrerythraea 34H, a Monomeric Cold Active Enzyme with Local Flexibility Around the Active Site and High Overall Stability.
Leiros, H.-K.S., Pey, A.L., Innselset, M., Moe, E., Leiros, I., Steen, I.H., Martinez, A.(2007) J Biological Chem 282: 21973
- PubMed: 17537732
- DOI: https://doi.org/10.1074/jbc.M610174200
- Primary Citation of Related Structures:
2V27, 2V28 - PubMed Abstract:
The characteristic of cold-adapted enzymes, high catalytic efficiency at low temperatures, is often associated with low thermostability and high flexibility. In this context, we analyzed the catalytic properties and solved the crystal structure of phenylalanine hydroxylase from the psychrophilic bacterium Colwellia psychrerythraea 34H (CpPAH). CpPAH displays highest activity with tetrahydrobiopterin (BH(4)) as cofactor and at 25 degrees C (15 degrees C above the optimal growth temperature). Although the enzyme is monomeric with a single L-Phe-binding site, the substrate binds cooperatively. In comparison with PAH from mesophilic bacteria and mammalian organisms, CpPAH shows elevated [S(0.5)](L-Phe) (= 1.1 +/- 0.1 mm) and K(m)(BH(4))(= 0.3 +/- 0.1 mm), as well as high catalytic efficiency at 10 degrees C. However, the half-inactivation and denaturation temperature is only slightly lowered (T(m) approximately 52 degrees C; where T(m) is half-denaturation temperature), in contrast to other cold-adapted enzymes. The crystal structure shows regions of local flexibility close to the highly solvent accessible binding sites for BH(4) (Gly(87)/Phe(88)/Gly(89)) and l-Phe (Tyr(114)-Pro(118)). Normal mode and COREX analysis also detect these and other areas with high flexibility. Greater mobility around the active site and disrupted hydrogen bonding abilities for the cofactor appear to represent cold-adaptive properties that do not markedly affect the thermostability of CpPAH.
- Norwegian Structural Biology Centre (NorStruct), Department of Chemistry, University of Tromsø, Tromsø.
Organizational Affiliation: