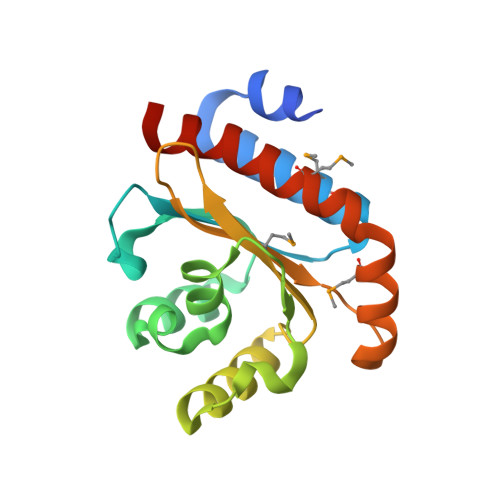
Molecular Insights Into Quorum Sensing in the Human Pathogen Pseudomonas Aeruginosa from the Structure of the Virulence Regulator Lasr Bound to its Autoinducer.
Bottomley, M.J., Muraglia, E., Bazzo, R., Carfi, A.(2007) J Biological Chem 282: 13592
- PubMed: 17363368 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M700556200
- Primary Citation Related Structures:
2UV0 - PubMed Abstract:
Many Gram-negative bacteria communicate via molecules called autoinducers to coordinate the activities of their populations. Such communication is termed quorum sensing and can regulate pathogenic virulence factor production and antimicrobial resistance. The quorum sensing system of Pseudomonas aeruginosa is currently the most intensively researched, because this bacterium is an opportunistic human pathogen annually responsible for the death of thousands of cystic fibrosis sufferers and many other immunocompromised individuals. Quorum sensing inhibitors can attenuate the pathogenicity of P. aeruginosa. Here we present the crystal structure of the P. aeruginosa LasR ligand-binding domain bound to its autoinducer 3-oxo-C(12)-acylhomoserine lactone. The structure is a symmetrical dimer, with each monomer exhibiting an alpha-beta-alpha fold similar to the TraR and SdiA quorum sensing proteins of Agrobacterium tumefaciens and Escherichia coli. The structure was determined up to 1.8-A resolution and reveals the atomic interactions between LasR and its autoinducer. The monomer structures of LasR, TraR, and SdiA are comparable but display differences in their quaternary organization. Inspection of their binding sites shows some unexpected variations resulting in quite different conformations of their bound autoinducers. We modeled interactions between LasR and various quorum sensing inhibitors, yielding insight into their possible mechanisms of action. The structure also provides a platform for the optimization, or de novo design, of quorum sensing inhibitors.
- Istituto di Ricerche di Biologia Molecolare, Via Pontina Km 30.600, 00040 Pomezia, Rome, Italy. matthew_bottomley@merck.com
Organizational Affiliation: