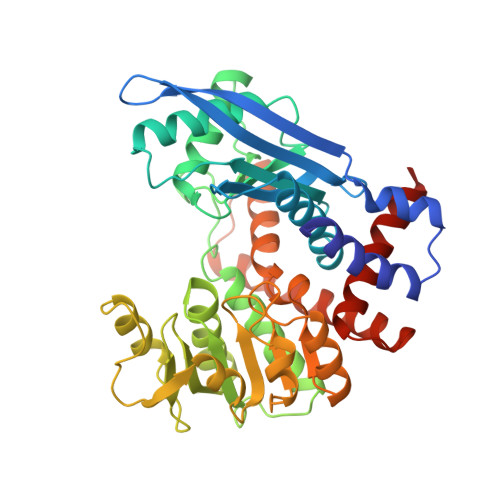
Engineering activity and stability of Thermotoga maritima glutamate dehydrogenase. II: construction of a 16-residue ion-pair network at the subunit interface.
Lebbink, J.H., Knapp, S., van der Oost, J., Rice, D., Ladenstein, R., de Vos, W.M.(1999) J Mol Biology 289: 357-369
- PubMed: 10366510
- DOI: https://doi.org/10.1006/jmbi.1999.2779
- Primary Citation Related Structures:
2TMG - PubMed Abstract:
The role of an 18-residue ion-pair network, that is present in the glutamate dehydrogenase from the hyperthermophilic archaeon Pyrococcus furiosus, in conferring stability to other, less stable homologous enzymes, has been studied by introducing four new charged amino acid residues into the subunit interface of glutamate dehydrogenase from the hyperthermophilic bacterium Thermotoga maritima. These two GDHs are 55 % identical in amino acid sequence, differ greatly in thermo-activity and stability and derive from microbes with different phylogenetic positions. Amino acid substitutions were introduced as single mutations as well as in several combinations. Elucidation of the crystal structure of the quadruple mutant S128R/T158E/N117R/S160E T. maritima glutamate dehydrogenase showed that all anticipated ion-pairs are formed and that a 16-residue ion-pair network is present. Enlargement of existing networks by single amino acid substitutions unexpectedly resulted in a decrease in resistance towards thermal inactivation and thermal denaturation. However, combination of destabilizing single mutations in most cases restored stability, indicating the need for balanced charges at subunit interfaces and high cooperativity between the different members of the network. Combination of the three destabilizing mutations in triple mutant S128R/T158E/N117R resulted in an enzyme with a 30 minutes longer half-life of inactivation at 85 degrees C, a 3 degrees C higher temperature optimum for catalysis, and a 0.5 degrees C higher apparent melting temperature than that of wild-type glutamate dehydrogenase. These findings confirm the hypothesis that large ion-pair networks do indeed stabilize enzymes from hyperthermophilic organisms.
- Laboratory of Microbiology, Wageningen Agricultural University, Hesselink van Suchtelenweg 4, CT Wageningen, NL-6703, The Netherlands. willem.devos@algemeen.micr.wau.nl
Organizational Affiliation: