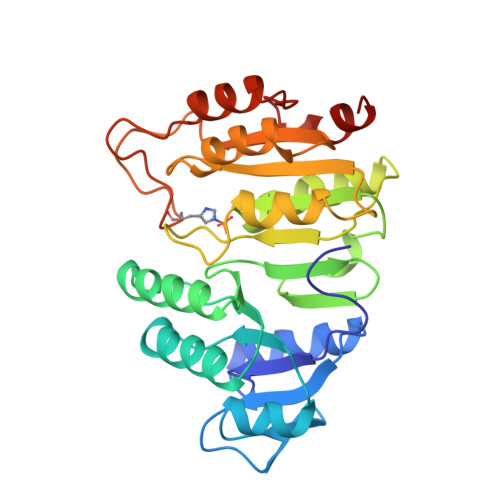
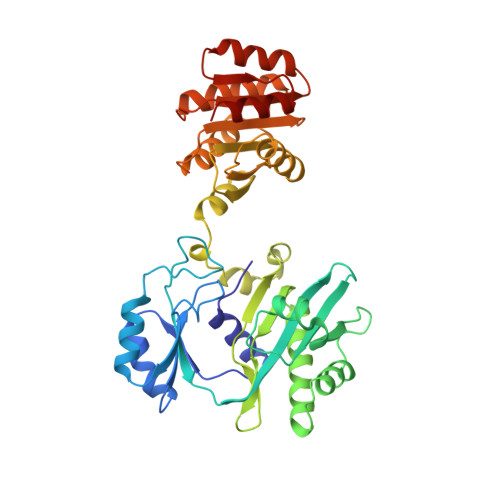
A detailed structural description of Escherichia coli succinyl-CoA synthetase.
Fraser, M.E., James, M.N., Bridger, W.A., Wolodko, W.T.(1999) J Mol Biology 285: 1633-1653
- PubMed: 9917402 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1998.2324
- Primary Citation Related Structures:
2SCU - PubMed Abstract:
Succinyl-CoA synthetase (SCS) carries out the substrate-level phosphorylation of GDP or ADP in the citric acid cycle. A molecular model of the enzyme from Escherichia coli, crystallized in the presence of CoA, has been refined against data collected to 2.3 A resolution. The crystals are of space group P4322, having unit cell dimensions a=b=98.68 A, c=403.76 A and the data set includes the data measured from 23 crystals. E. coli SCS is an (alphabeta)2-tetramer; there are two copies of each subunit in the asymmetric unit of the crystals. The crystal packing leaves two choices for which pair of alphabeta-dimers form the physiologically relevant tetramer. The copies of the alphabeta-dimer are similar, each having one active site where the phosphorylated histidine residue and the thiol group of CoA are found. CoA is bound in an extended conformation to the nucleotide-binding motif in the N-terminal domain of the alpha-subunit. The phosphoryl group of the phosphorylated histidine residue is positioned at the amino termini of two alpha-helices, one from the C-terminal domain of the alpha-subunit and the other from the C-terminal domain of the beta-subunit. These two domains have similar topologies, despite only 14 % sequence identity. By analogy to other nucleotide-binding proteins, the binding site for the nucleotide may reside in the N-terminal domain of the beta-subunit. If this is so, the catalytic histidine residue would have to move about 35 A to react with the nucleotide.
- Department of Biochemistry, University of Alberta, Canada.
Organizational Affiliation: