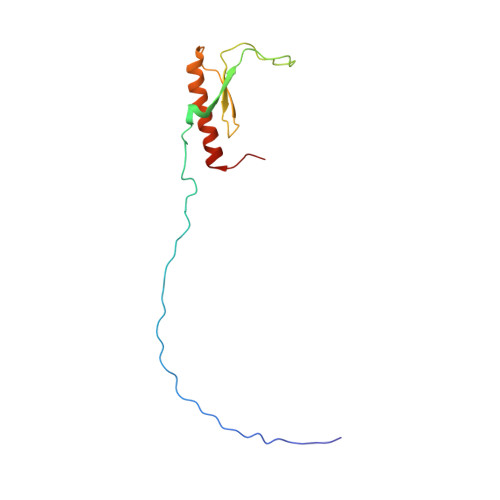
Solution structure and siRNA-mediated knockdown analysis of the mitochondrial disease-related protein C12orf65.
Kogure, H., Hikawa, Y., Hagihara, M., Tochio, N., Koshiba, S., Inoue, Y., Guntert, P., Kigawa, T., Yokoyama, S., Nameki, N.(2012) Proteins
- PubMed: 22821833 Search on PubMed
- DOI: https://doi.org/10.1002/prot.24152
- Primary Citation Related Structures:
2RSM - PubMed Abstract:
Loss of function of the c12orf65 gene causes a mitochondrial translation defect, leading to encephalomyopathy. The C12orf65 protein is thought to play a role similar to that of ICT1 in rescuing stalled mitoribosomes during translation. Both proteins belong to a family of Class I peptide release factors (RFs), all characterized by the presence of a GGQ motif. Here, we determined the solution structure of the GGQ-containing domain (GGQ domain) of C12orf65 from mouse by NMR spectroscopy, and examined the effect of siRNA-mediated knockdown of C12orf65 on mitochondria in HeLa cells using flow cytometry. The GGQ domain, comprising residues 60-124 of the 184-residue full-length protein, forms a structure with a 3(10) -β1-β2-β3-α1 topology that resembles the GGQ domain structure of RF more closely than that of ICT1. Thus, the GGQ domain structures of this protein family can be divided into two types, depending on the region linking β2 and β3; the C12orf65/RF type having a 6-residue π-HB turn and the ICT1 type having an α-helix. Knockdown of C12orf65 resulted in increased ROS production and apoptosis, leading to inhibition of cell proliferation. Substantial changes in mitochondrial membrane potential and mass in the C12orf65-knockdown cells were observed compared with the control cells. These results indicate that the function of C12orf65 is essential for cell vitality and mitochondrial function. Although similar effects were observed in ICT1-downregulated cells, there were significant differences in the range and pattern of the effects between C12orf65- and ICT1-knockdown cells, suggesting different roles of C12orf65 and ICT1 in rescuing stalled mitoribosomes.
- Department of Chemistry and Chemical Biology, Graduate School of Engineering, Gunma University, Kiryu, Gunma 376-8515, Japan.
Organizational Affiliation: