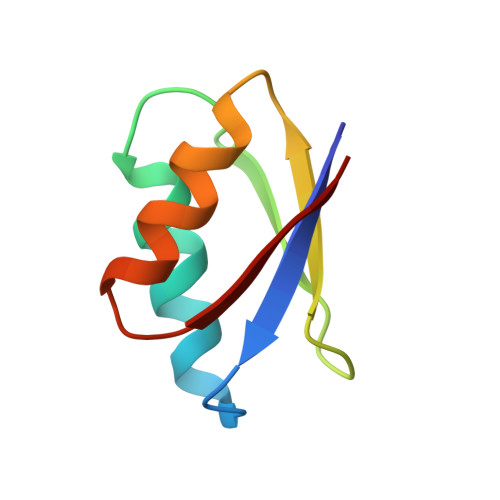
Protein structure determination in living cells by in-cell NMR spectroscopy
Sakakibara, D., Sasaki, A., Ikeya, T., Hamatsu, J., Hanashima, T., Mishima, M., Yoshimasu, M., Hayashi, N., Mikawa, T., Walchli, M., Smith, B.O., Shirakawa, M., Guntert, P., Ito, Y.(2009) Nature 458: 102-105
- PubMed: 19262674 Search on PubMed
- DOI: https://doi.org/10.1038/nature07814
- Primary Citation Related Structures:
2ROE, 2ROG - PubMed Abstract:
Investigating proteins 'at work' in a living environment at atomic resolution is a major goal of molecular biology, which has not been achieved even though methods for the three-dimensional (3D) structure determination of purified proteins in single crystals or in solution are widely used. Recent developments in NMR hardware and methodology have enabled the measurement of high-resolution heteronuclear multi-dimensional NMR spectra of macromolecules in living cells (in-cell NMR). Various intracellular events such as conformational changes, dynamics and binding events have been investigated by this method. However, the low sensitivity and the short lifetime of the samples have so far prevented the acquisition of sufficient structural information to determine protein structures by in-cell NMR. Here we show the first, to our knowledge, 3D protein structure calculated exclusively on the basis of information obtained in living cells. The structure of the putative heavy-metal binding protein TTHA1718 from Thermus thermophilus HB8 overexpressed in Escherichia coli cells was solved by in-cell NMR. Rapid measurement of the 3D NMR spectra by nonlinear sampling of the indirectly acquired dimensions was used to overcome problems caused by the instability and low sensitivity of living E. coli samples. Almost all of the expected backbone NMR resonances and most of the side-chain NMR resonances were observed and assigned, enabling high quality (0.96 ångström backbone root mean squared deviation) structures to be calculated that are very similar to the in vitro structure of TTHA1718 determined independently. The in-cell NMR approach can thus provide accurate high-resolution structures of proteins in living environments.
- Department of Chemistry, Tokyo Metropolitan University, Hachioji, Tokyo 192-0397, Japan.
Organizational Affiliation: