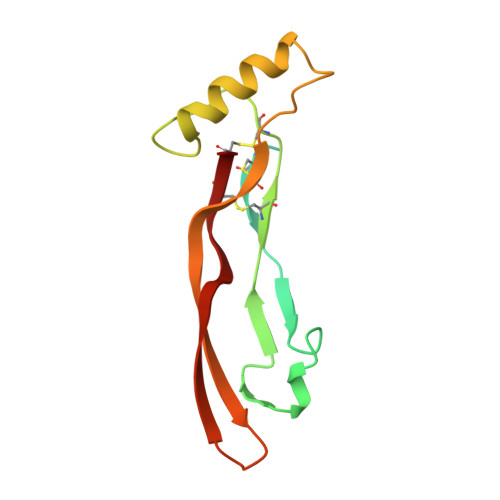
Type I receptor binding of bone morphogenetic protein 6 is dependent on N-glycosylation of the ligand.
Saremba, S., Nickel, J., Seher, A., Kotzsch, A., Sebald, W., Mueller, T.D.(2007) FEBS J 275: 172-183
- PubMed: 18070108 Search on PubMed
- DOI: https://doi.org/10.1111/j.1742-4658.2007.06187.x
- Primary Citation Related Structures:
2R52, 2R53 - PubMed Abstract:
Bone morphogenetic proteins (BMPs), together with transforming growth factor (TGF)-beta and activins/inhibins, constitute the TGF-beta superfamily of ligands. This superfamily is formed by more than 30 structurally related secreted proteins. The crystal structure of human BMP-6 was determined to a resolution of 2.1 A; the overall structure is similar to that of other TGF-beta superfamily ligands, e.g. BMP-7. The asymmetric unit contains the full dimeric BMP-6, indicating possible asymmetry between the two monomeric subunits. Indeed, the conformation of several loops differs between both monomers. In particular, the prehelix loop, which plays a crucial role in the type I receptor interactions of BMP-2, adopts two rather different conformations in BMP-6, indicating possible dynamic flexibility of the prehelix loop in its unbound conformation. Flexibility of this loop segment has been discussed as an important feature required for promiscuous binding of different type I receptors to BMPs. Further studies investigating the interaction of BMP-6 with different ectodomains of type I receptors revealed that N-glycosylation at Asn73 of BMP-6 in the wrist epitope is crucial for recognition by the activin receptor type I. In the absence of the carbohydrate moiety, activin receptor type I-mediated signaling of BMP-6 is totally diminished. Thus, flexibility within the binding epitope of BMP-6 and an unusual recognition motif, i.e. an N-glycosylation motif, possibly play an important role in type I receptor specificity of BMP-6.
- Lehrstuhl für Physiologische Chemie II, Biozentrum der Universität Würzburg, Germany.
Organizational Affiliation: