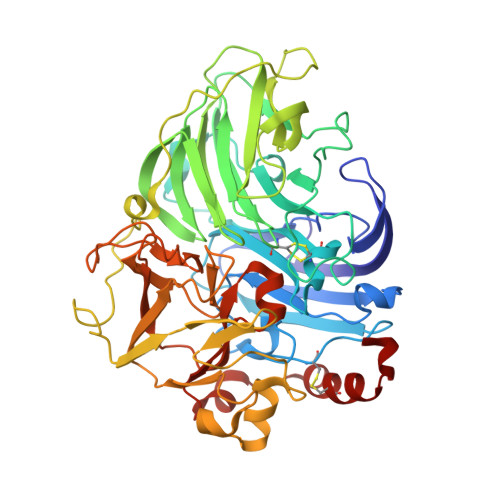
Crystal structure of a blue laccase from Lentinus tigrinus: evidences for intermediates in the molecular oxygen reductive splitting by multicopper oxidases.
Ferraroni, M., Myasoedova, N.M., Schmatchenko, V., Leontievsky, A.A., Golovleva, L.A., Scozzafava, A., Briganti, F.(2007) BMC Struct Biol 7: 60-60
- PubMed: 17897461 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1186/1472-6807-7-60
- Primary Citation Related Structures:
2QT6 - PubMed Abstract:
Laccases belong to multicopper oxidases, a widespread class of enzymes implicated in many oxidative functions in pathogenesis, immunogenesis and morphogenesis of organisms and in the metabolic turnover of complex organic substances. They catalyze the coupling between the four one-electron oxidations of a broad range of substrates with the four-electron reduction of dioxygen to water. These catalytic processes are made possible by the contemporaneous presence of at least four copper ion sites, classified according to their spectroscopic properties: one type 1 (T1) site where the electrons from the reducing substrates are accepted, one type 2 (T2), and a coupled binuclear type 3 pair (T3) which are assembled in a T2/T3 trinuclear cluster where the electrons are transferred to perform the O2 reduction to H2O. The structure of a laccase from the white-rot fungus Lentinus (Panus) tigrinus, a glycoenzyme involved in lignin biodegradation, was solved at 1.5 A. It reveals a asymmetric unit containing two laccase molecules (A and B). The progressive reduction of the copper ions centers obtained by the long-term exposure of the crystals to the high-intensity X-ray synchrotron beam radiation under aerobic conditions and high pH allowed us to detect two sequential intermediates in the molecular oxygen reduction pathway: the "peroxide" and the "native" intermediates, previously hypothesized through spectroscopic, kinetic and molecular mechanics studies. Specifically the electron-density maps revealed the presence of an end-on bridging, micro-eta 1:eta 1 peroxide ion between the two T3 coppers in molecule B, result of a two-electrons reduction, whereas in molecule A an oxo ion bridging the three coppers of the T2/T3 cluster (micro3-oxo bridge) together with an hydroxide ion externally bridging the two T3 copper ions, products of the four-electrons reduction of molecular oxygen, were best modelled. This is the first structure of a multicopper oxidase which allowed the detection of two intermediates in the molecular oxygen reduction and splitting. The observed features allow to positively substantiate an accurate mechanism of dioxygen reduction catalyzed by multicopper oxidases providing general insights into the reductive cleavage of the O-O bonds, a leading problem in many areas of biology.
- Department of Chemistry University of Florence, Via della Lastruccia 3, Sesto Fiorentino, 50019 Florence, Italy. mferraroni@unifi.it
Organizational Affiliation: