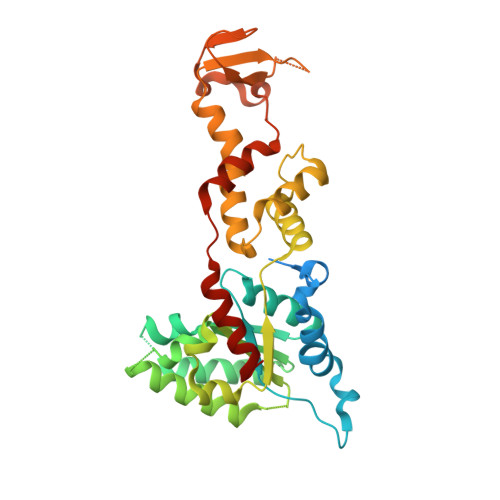
Structural characterization of the ATPase reaction cycle of endosomal AAA protein Vps4.
Xiao, J., Xia, H., Yoshino-Koh, K., Zhou, J., Xu, Z.(2007) J Mol Biology 374: 655-670
- PubMed: 17949747 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2007.09.067
- Primary Citation Related Structures:
2QP9, 2QPA - PubMed Abstract:
The multivesicular body (MVB) pathway functions in multiple cellular processes including cell surface receptor down-regulation and viral budding from host cells. An important step in the MVB pathway is the correct sorting of cargo molecules, which requires the assembly and disassembly of endosomal sorting complexes required for transport (ESCRTs) on the endosomal membrane. Disassembly of the ESCRTs is catalyzed by ATPase associated with various cellular activities (AAA) protein Vps4. Vps4 contains a single AAA domain and undergoes ATP-dependent quaternary structural change to disassemble the ESCRTs. Structural and biochemical analyses of the Vps4 ATPase reaction cycle are reported here. Crystal structures of Saccharomyces cerevisiae Vps4 in both the nucleotide-free form and the ADP-bound form provide the first structural view illustrating how nucleotide binding might induce conformational changes within Vps4 that lead to oligomerization and binding to its substrate ESCRT-III subunits. In contrast to previous models, characterization of the Vps4 structure now supports a model where the ground state of Vps4 in the ATPase reaction cycle is predominantly a monomer and the activated state is a dodecamer. Comparison with a previously reported human VPS4B structure suggests that Vps4 functions in the MVB pathway via a highly conserved mechanism supported by similar protein-protein interactions during its ATPase reaction cycle.
- Life Sciences Institute and Department of Biological Chemistry, Medical School, University of Michigan, Ann Arbor, MI 48109, USA.
Organizational Affiliation: